LncRNA Microarrays
Related Products
Related Services
Related Reviews
LncRNA Array Service
Arraystar LncRNA Arrays are designed to systematically profile long non-coding RNAs (lncRNAs) along with the entire set of protein-coding mRNAs. They are the best tool for profiling LncRNAs, overcoming many limitations of RNA-seq for lncRNAs often at low abundance.
To incorporate rapid scientific advances and new data, Arraystar has newly released LncRNA Array versions updated to Human V5.0, Mouse V4.0, and Rat V3.0. These new arrays provide the most current, robust, full length LncRNA collections, complete with major upgrades in scientific annotation and analyses.
Benefits
• Most sensitive and best technology for lncRNA profiling, superior to RNA-seq. Learn more>>
• Comprehensive and robust full-length* LncRNA collection curated from the all major latest databases and landmark publications. Learn more>>
• Systematic and specialized lncRNA annotation, including genomic context, epigenomic context*, completeness*, subcellular localization**, miRNA recognition… Learn more>>
• An invaluable tool for LncRNA research with hundreds of high impact publications. Learn more>>
*Applied to Human V5.0 ** Applied to Human V5.0 and Mouse V4.0
Watch Video> How to Study LncRNA Expression and Modifications?
Watch Video> The Latest Development of LncRNA Profiling Technology
| Service Name | Catalog No | Description | Format | Price |
|---|---|---|---|---|
| Human V5.0 LncRNA Array Service | AS-S-LNC-H | 39,317 LncRNAs and 21,174 mRNAs | 8*60K | |
| Mouse V4.0 LncRNA Array Service | AS-S-LNC-M | 37,949 LncRNAs and 22,692 mRNAs | 8*60K | |
| Rat V3.0 LncRNA Array Service | AS-S-LNC-R | 10,333 LncRNAs and 28,287 mRNAs | 4*44K |
More sensitive and better technology than RNA-seq for lncRNA profiling.
Long non-coding RNAs (LncRNAs) often express and function at low abundance, buried in other classes of abundant RNAs [1]. There are serious limitations of RNA-seq for lncRNA profiling.
• Poor quantification of lncRNAs due to the low abundance and the lack of completeness in lncRNA annotation
• Ambiguous detection of LncRNA isoforms caused by the weak splice profile and the missing connectivity information
• Systematic and functional lncRNA annotation database publicly unavailable for RNA seq analysis
Comprehensive, robust, and most up-to-date LncRNA array contents.
Unlike protein coding genes, publicly available lncRNAs are often scantily annotated, partial in scope and scattered in collection. Arraystar maintains high quality proprietary lncRNA transcriptome databases that extensively collect lncRNAs through all major public databases and repositories, knowledge-based mining of scientific publications, and our lncRNA discovery pipelines:
Human V5.0: FANTOM5 CAT (v1), GENCODE (v29), RefSeq (Updated to 2018.11), BIGTranscriptome (v1), knownGene (Updated to 2018.11), LncRNAdb, LncRNAWiki, RNAdb, NRED, CLS FL, NONCODE (v5), MiTranscriptome (v2), lncRNA discovery pipeline from more than 47 Tb RNA-seq data. A total of 39,317 lncRNAs in Gold Standard and Reliable LncRNA tiers. Learn more>>
Mouse V4.0: GENCODE(VM19), RefSeq(Updated to 2018.11), KnownGene(Updated to 2018.11), GenBank.
Rat V3.0: Ensembl (94.rn6), RefSeq(Updated to 2019.01.03).
Systematic lncRNA annotations and analyses incorporating the advances in lncRNA research.
Complete with genomic information, epigenomic context, subclassification, transcript model completeness, subcellular localization, miRNA recognition, conservation, tissue/cell specificity, and small peptide coding potentials, biological process, and disease association, for gaining insights into the complex lncRNA biology.
Both noncoding lncRNA and protein-coding mRNA expression datasets on the same array to allow their co-expressional and correlational studies.
Human LncRNA Expression Array V5.0
| Total number of distinct probes | 60,491 |
| Probe length | 60 nt |
| Probe site | mRNA and lncRNA: Specific exon or splice junction sequence along the entire length of transcript. |
| Probe specificity | Transcript-specific |
| Protein coding mRNAs | 21,174 |
| LncRNAs | 39,317 (8,393 Gold Standard LncRNAs and 30,924 Reliable LncRNAs) |
| mRNA sources | Refseq, UCSC, GENCODE, FANTOM5 CAT |
| LncRNA sources | Arraystar LncRNA collection pipelines: lncRNAs from all major databases and literatures up to 2018. “Canonical” or “longest” priority assigned to the transcript for each lncRNA gene. External Databases (current in 2018): FANTOM5 CAT (v1), GENCODE (v29), RefSeq (Updated to 2018.11), BIGTranscriptome (v1), knownGene (Updated to 2018.11), LncRNAdb, LncRNAWiki, RNAdb, NRED, CLS FL, NONCODE (v5), MiTranscriptome (v2) Literatures: Scientific publications up to 2018 |
| Array Format | 8 × 60 K |
Mouse LncRNA Expression Array V4.0
| Total number of distinct probes | 60,641 |
| Probe length | 60 nt |
| Probe site | mRNA and lncRNA: Specific exon or splice junction sequence along the entire length of transcript. |
| Probe specificity | Transcript-specific |
| Protein coding mRNAs | 22,692 |
| LncRNAs | 37,949 |
| mRNA sources | Refseq, Known Gene, GENCODE |
| LncRNA sources | Arraystar LncRNA collection pipelines: lncRNAs from all major databases and literatures up to 2018. “Canonical” or “longest” priority assigned to the transcript for each lncRNA gene. External Databases (current in 2018): GENECODE(VM19), RefSeq, KnownGene, GenBank Literatures: Scientific publications up to 2018 |
| Array Format | 8 × 60 K |
Rat LncRNA Expression Array V3.0
| Total number of distinct probes | 38,620 |
| Probe length | 60 nt |
| Probe site | mRNA and lncRNA: Specific exon or splice junction sequence along the entire length of transcript. |
| Probe specificity | Transcript-specific |
| Protein coding mRNAs | 28,287 |
| LncRNAs | 10,333 |
| mRNA sources | Refseq, Ensembl |
| LncRNA sources | Arraystar LncRNA collection pipelines: lncRNAs from all major databases and literatures up to 2018. “Canonical” or “longest” priority assigned to the transcript for each lncRNA gene. External Databases (current in 2018): Refseq, Ensembl Literatures: Scientific publications up to 2018 |
| Array Format | 4 × 44 K |
Arraystar’s specially-designed LncRNA Arrays are available only through our LncRNA Array service. We provide full-service LncRNA Array profiling, from sample preparation to in-depth data analysis. Our step-by-step quality controls are designed to ensure you get the most reliable results. Just send us your samples, and we’ll do the rest!
Please refer to Sample Submission for details in how to get your project started.
• RNA isolation (Optional)
• RNA QC
• cDNA synthesis
• Target preparation by labeling with Cy3
• Array hybridization, washing, and scanning
• Data extraction, analysis and summarization
Arraystar’s experienced scientists have intimate knowledge of the LncRNA array platform and are experts in the analysis and interpretation of LncRNA profiling data. This familiarity enables us to employ the most robust methods for normalization and data analysis including subgroup classification.
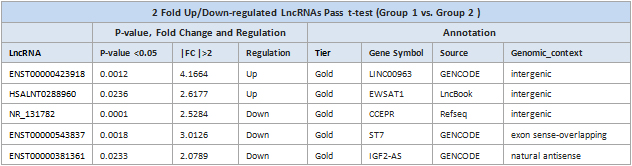
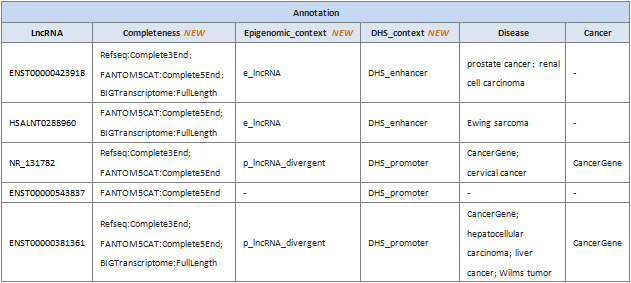
Table 1. Differentially expressed LncRNAs with systematic lncRNA annotations. Differentially expressed LncRNAs (Fold change>2, p-value<0.05) in Group1 vs Group2. In column “Regulation”, “Up” indicates up-regulated, “Down” indicates down-regulated.
Deep data mining and advanced analyses (Fig. 1 and 2, for example) are available to carry you further with the wealth of information generated by the microarrays.
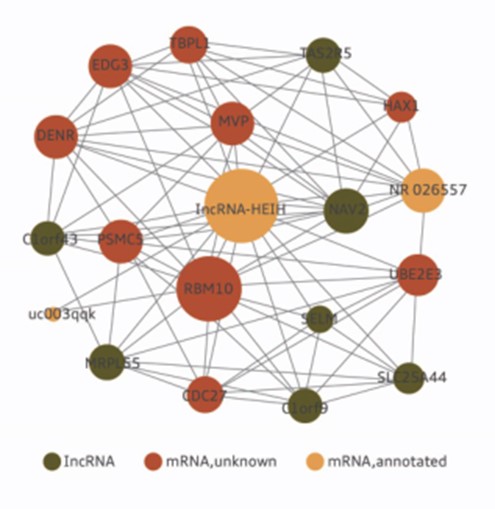
Figure 1. Coding-noncoding interaction Co-expression (CNC) subnetwork of an lncRNA-HEIH in hepatocellular carcinoma.
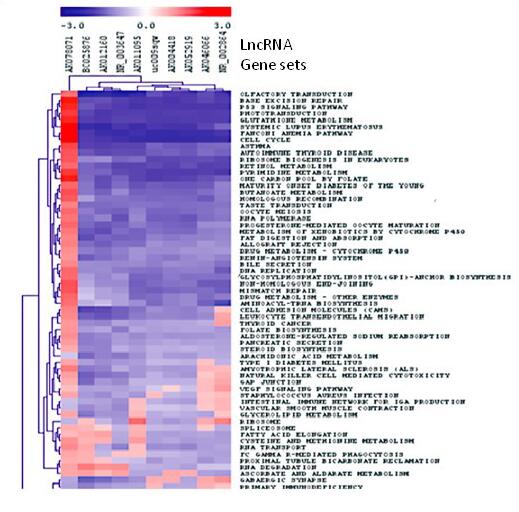
Figure 2. Lnc-GSET to identify enrichment of lncRNAs in biological functions.
Targeting Ischemic Myocardium: Nanoparticles Loaded with Long Noncoding RNA AK156373 siRNA Alleviate Myocardial Infarction. Gao M,et al. ACS Nano, 2025
Identification of biomarkers of shrinkage modes after neoadjuvant therapy in HER-2 positive breast cancer. Bi Z, et al. International Journal of Surgery, 2025
A novel lncRNA ROPM-mediated lipid metabolism governs breast cancer stem cell properties. Liu S, et al. Journal of Hematology & Oncology, 2024
LncRNA evf-2 Exacerbates Podocyte Injury in Diabetic Nephropathy by Inducing Cell Cycle Re-entry and Inflammation Through Distinct Mechanisms Triggered by hnRNPU. Zhang C, et al. Advanced Science, 2024
RRFERV stabilizes TEAD1 expression to mediate nasopharyngeal cancer radiation resistance rendering tumor cells vulnerable to ferroptosis. Xu Q, et al. International Journal of Surgery, 2024
LncRNA AL139294.1 can be transported by extracellular vesicles to promote the oncogenic behaviour of recipient cells through activation of the Wnt and NF-kB2 pathways in non-small-cell lung cancer. Ma X, et al. Journal of Experimental & Clinical Cancer Research, 2024
LIN28B induced PCAT5 promotes endometrial cancer progression and glycolysis via IGF2BP3 deubiquitination. Wang B, et al. Cell Death & Disease, 2024
Integrative analysis discovers Imidurea as dual multitargeted inhibitor of CD69, CD40, SHP2, lysozyme, GATA3, cCBL, and S-cysteinase from SARS-CoV-2 and M. tuberculosis. Ahmad S, et al. International journal of biological macromolecules, 2024
High-density lipoprotein regulates angiogenesis by long non-coding RNA HDRACA. Mo Z W,et al. Signal Transduction and Targeted Therapy, 2023
LncRNA PSMB8-AS1 Instigates Vascular Inflammation to Aggravate Atherosclerosis. Li S, et al. Circulation Research, 2023
FAR591 promotes the pathogenesis and progression of SONFH by regulating Fos expression to mediate the apoptosis of bone microvascular endothelial cells. Zhang F, et al. Bone Research, 2023
LncRNA NIPA1-SO confers atherosclerotic protection by suppressing the transmembrane protein NIPA1. Jiang M,et al. Journal of Advanced Research, 2023
A lncRNA signature associated with tumor immune heterogeneity predicts distant metastasis in locoregionally advanced nasopharyngeal carcinoma. Liang Y L, et al, Nature Communications, 2022
The long noncoding RNA glycoLINC assembles a lower glycolytic metabolon to promote glycolysis. Zhu Y, et al. Molecular Cell, 2022
Mmu-LncRNA 121686/hsa-lncRNA 520657 induced by METTL3 drive the progression of AKI by targeting miR-328-5p /HtrA3 signaling axis. Pan J,et al. Molecular Therapy, 2022
Long noncoding RNA ERLR mediates epithelial-mesenchymal transition of retinal pigment epithelial cells and promotes experimental proliferative vitreoretinopathy. Yang S, et al. Cell Death & Differentiation, 2021
Defining the adult neural stem cell niche proteome identifies key regulators of adult neurogenesis. Kjell J, et al. Cell stem cell, 2020
lncRNA JPX/miR-33a-5p/Twist1 axis regulates tumorigenesis and metastasis of lung cancer by activating Wnt/ß-catenin signaling. Pan J, et al. Molecular Cancer, 2020
H. pylori infection alters repair of DNA double-strand breaks via SNHG17. Han T, et al. The Journal of Clinical Investigation, 2020
Fibrogenic Activity of MECP2 is Regulated by Phosphorylation in Hepatic Stellate Cells. Moran-Salvador E, et al. Gastroenterology, 2019
A long noncoding RNA cluster-based genomic locus maintains proper development and visual function. Wang F,et al. Nucleic Acids Research, 2019
LncRNA PCAT1 activates AKT and NF-kB signaling in castration-resistant prostate cancer by regulating the PHLPP/FKBP51/IKKa complex. Shang Z, et al. Nucleic Acids Research, 2019
Loss of RNA-binding protein GRSF1 activates mTOR to elicit a proinflammatory transcriptional program. Noh J H, et al. Nucleic Acids Research, 2019
NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Huang D, et al. Nature immunology, 2018
Long noncoding RNA lnc-TSI inhibits renal fibrogenesis by negatively regulating the TGF-ß/Smad3 pathway. Wang P, et al. Science translational medicine, 2018
GUARDIN is a p53-responsive long non-coding RNA that is essential for genomic stability. Hu W L, et al. Nature Cell Biology, 2018
LncRNA-FEZF1-AS1 promotes tumor proliferation and metastasis in colorectal cancer by regulating PKM2 signaling. Bian Z, et al. Clinical Cancer Research, 2018
A novel lncRNA GClnc1 promotes gastric carcinogenesis and may act as a modular scaffold of WDR5 and KAT2A complexes to specify the histone modification pattern. Sun T T, et al. Cancer discovery, 2016
The LINK-A lncRNA interacts with PtdIns (3, 4, 5) P3 to hyperactivate AKT and confer resistance to AKT inhibitors. Lin A, et al. Nature Cell Biology, 2017
Integrative Transcriptome Analyses of Metabolic Responses in Mice Define Pivotal LncRNA Metabolic Regulators. Ling Yang, et al. Cell Metabolism, 2016
Wnt signalling modulates transcribed-ultraconserved regions in hepatobiliary cancers. Carotenuto P, et al. Gut, 2016
Differential transcriptome expression in human nucleus accumbens as a function of loneliness. Canli T, et al. Molecular Psychiatry, 2016
Long noncoding RNA Tug1 regulates mitochondrial bioenergetics in diabetic nephropathy. Long J, et al. The Journal of Clinical Investigation, 2016
The senescence-associated secretory phenotype is potentiated by feedforward regulatory mechanisms involving Zscan4 and TAK1. Zhang B, et al. Nature communications, 2018
Non-coding RNAs participate in the regulatory network of CLDN4 via ceRNA mediated miRNA evasion. Yong-xi Song,et al. Nature communications, 2017
Long Non-coding RNA DILC Represses Self-renewal of Liver Cancer Stem Cells via Inhibiting Autocrine IL-6/STAT3 Axis. X Wang, et al. Journal of Hepatology, 2016
