The Arraystar LncRNA microarray package includes systematic and detailed long non-coding RNA (lncRNA) annotations, subclassification, and analyses to gain insight into the complex biological functions of the lncRNAs. LncRNAs with reported biological processes or associated with human diseases are researched, annotated and cross referenced. This rich source of information helps to unravel functional roles and molecular mechanisms of the LncRNAs.
Genomic context. LncRNAs are systematically classified based on their genomic relationships with the nearest protein coding genes into Intergenic (LincRNA), Intronic, Bidirectional, Sense-overlapping, Antisense and Pseudogene LncRNAs (Fig. 1). These subclasses help dissect various cis- or trans-regulatory functions on the target genes transcriptionally or post-transcriptionally (Fig. 2).
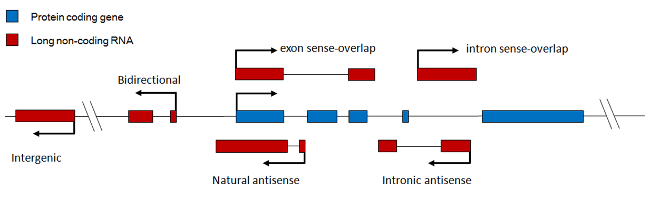
Figure 1. LncRNA classification and subgroups in various genomic context with the protein coding gene.
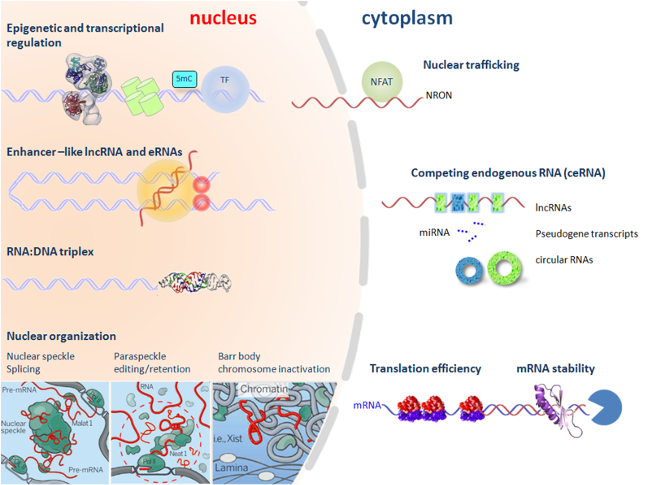
Figure 2. LncRNAs may regulate gene expression by various mechanisms, such as recruiting chromatin modifiers/remodelers to epigenetically regulate gene expression; by enhancer RNAs; by nuclear substructures; by nuclear-cytoplasmic transport; by competing endogenous RNAs via miRNAs; or by mRNA stability and translation; in cis or in trans, at transcription or post-transcriptional levels.
Epigenomic context*. lncRNAs can be transcribed in and regulated by a promoter or enhancer region with characteristic promoter or enhancer epigenetic marks. Many active promoter and enhancer regions are themselves transcription units, capable of generating functionally active noncoding RNAs for these cis-regulatory DNA elements. The lncRNAs are thus classified into promoter-lncRNAs (p-lncRNA) and enhancer-lncRNAs (e-lncRNA) based on the epigenomic context (e.g. DNase I hypersensitive sites). The p-lncRNAs are further grouped into intergenic and divergent p-lncRNAs based on their genomic context (Fig. 3). p-lncRNAs are often positively correlated with transcription of their protein-coding genes under the same promoters. e-lncRNAs often trap TF proteins to the local sites, modify the local chromatin environment, and organize three-dimensional nuclear topology domains for correct activation of target gene program.
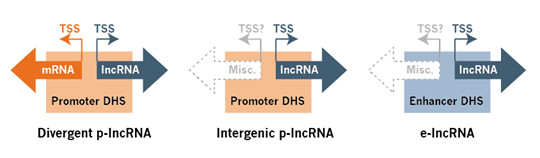
Figure 3. Promoter and enhancer lncRNA categories based on the epigenomic and genomic context. lncRNAs are classified into intergenic p-lncRNA, divergent p-lncRNA, e-lncRNA, and other, based on their TSS and DNase I hypersensitive sites (DHS) in the promoter (marked by H3K4me3), enhancer (marked by H3K4me1, H3K27ac and H3K9ac), or dyadic regulatory (enhancer-promoter alternating states) regions.
Completeness*. The sequence completeness of the lncRNA 5’ and 3’ ends are important for studying the lncRNAs. For example, accurate lncRNA transcription start sites (TSS) are particularly important for identifying the lncRNA promoters or designing CRISPR–Cas screen targeting sites. Here, the lncRNA completeness status is annotated in three categories: Complete 5’end, Complete 3’end, and Full length (complete both 5’ and 3’end).
Subcellular localization**. The molecular functions of lncRNAs are tightly coupled with their subcellular localization. For example, lncRNAs localized in the nucleus or chromatin often regulate the gene expression by epigenetic modification and transcription. LncRNAs in the cytoplasm are more likely involved in translation regulation or miRNA sponging such as competing endogenous RNAs (ceRNA).
miRNA recognition site. Predicted or experimentally identified microRNA sites on the lncRNAs are annotated to indicate potential post-transcriptional regulatory functions in the miRNA regulatory network, such as acting as competing endogenous RNAs (ceRNA).
Highly conserved lncRNAs. Certain lncRNA genes harbor ultraconserved regions (UCR) or ultraconserved non-coding elements (UCNEs) that do not vary in sequence across species, which imply these sequences being biologically indispensible. As many lncRNAs regulate target genes by cis-mechanism, human lncRNAs syntenic to orthologous lncRNAs in other species are also collected even with modest homology, as their genomic context with the neighboring target genes, rather than the sequence conservation, can be more relevant in gene regulation [1].
Tissue specific lncRNAs*. The function of a lncRNA can be directly or indirectly related to and indicated by the tissue or cell type in which it is specifically expressed [1]. In the lncRNA microarray, 6,059 cell lineage or cancer associated lncRNAs are annotated [2].
Disease-associated lncRNAs*. LncRNAs known to be associated with diseases, such as cataloged in LncRNADisease [3], are annotated for clinical and translational investigations.
Coding potential for small peptides*. Although most lncRNAs are noncoding, some lncRNAs can contain small open reading frames (smORFs) to encode small peptides as predicted or experimentally detected as cataloged in lncRNAWiki.
*Applied to Human V5.0 ** Applied to Human V5.0 and Mouse V4.0
Related Service
