Featured ncRNA Microarrays
Related Products
Related Services
Related Reviews
Small RNA Array Service
Arraystar Small RNA Array uses direct end-labeling and smart probe design microarray technologies to detect and quantify small RNAs including miRNA, pre-miRNA, tsRNAs, tRNAs, and snoRNAs simultaneously on the same array, providing vital expressional information to study small RNA regulation and biomarker applications.
Benefits
• Simultaneously profile the major small RNA classes: miRNA, pre-miRNA, tRNA, tsRNA, and snoRNA
• Raise the bar of small RNA profiling to high sensitivity, specificity and accuracy by direct end-labeling and smart probe design.
• Direct and simplified procedures to overcome biases from RNA modifications, RNA fold hindrance, reverse transcription blocks, PCR amplifications, and analysis inaccuracy in small RNA-seq. Learn more>>
• Required RNA sample amounts starting as low as 100 ng, opening up many research opportunities. Learn more>>
• Tolerant for RNA samples at lower qualities: e.g. degraded RNAs, serum/plasma/biofluid RNAs, FFPE RNAs. Learn more>>
Watch Video> Raising the Bar of Multi-transcriptomic Profiling of Small RNAs
| Service Name | Description | Format | Price |
|---|---|---|---|
| Human Small RNA Array Service | Profile human miRNA, pre-miRNA, tsRNA, tRNA&snoRNA | 8 x 15K | |
| Mouse Small RNA Array Service | Profile mouse miRNA, pre-miRNA, tsRNA, tRNA&snoRNA | 8 x 15K | |
| Rat Small RNA Array Service | Profile rat miRNA, pre-miRNA, tsRNA, tRNA&snoRNA | 8 x 15K |
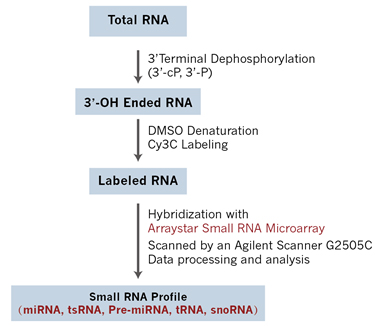
Figure 1. Arraystar Small RNA Array Profiling workflow. Total RNA is treated with T4 polynucleotide kinase (T4 PNK) to remove residual phosphate (P) and cyclic phosphate (cP) groups from the 3′ end to expose the 3-OH. The 3-OH ended small RNAs are denatured by DMSO and enzymatically labeled with Cy3 by ligation. The labeled RNAs are hybridized onto Arraystar Small RNA Expression Microarray for analysis of the small RNA expression profiles.
• Simultaneous profiling of the major small RNA classes
Arraystar Small RNA Array contents include miRNA, pre-miRNA, tsRNAs, tRNAs and snoRNAs. These small RNAs are of particular interest for biological and clinical sciences for their important gene regulation functions and great biomarker potentials. Profiling by small RNA-seq requires separate chemistries for sequencing library construction and analyses to cover small RNA classes with distinct biochemical properties. For example, miRNA-seq and tRNA-seq cannot be run together to profile both miRNA and tRNA biotypes. Now Arraystar Small RNA Array can cover all these small RNAs simultaneously on one chip in one experiment.
• Raise the bar of small RNA profiling for high sensitivity, specificity and accuracy
Arraystar Small RNA Array uses high affinity probe hybridization to achieve very high sensitivity even for small RNAs at low abundance. The smart probe design incorporates 5’-hairpin structure and normalized sequence targeting region to specifically distinguish small RNAs with only 1~2 nucleotide differences. Learn more>>
The direct and simplified procedures ensure unbiased and accurate quantification better than small RNA-seq or even qPCR.
The long standing challenges of small RNA profiling by RNA-seq have been RNA modifications, RNA fold structures, and complicated RNA-seq library preparation. In Arraystar Small RNA Array, small RNAs are simply ligated with Cy3 dye directly at their 3’-OH ends, which complete avoids RNA pretreatments to remove internal RNA modifications, abortive reverse transcription due to the modifications and RNA fold hindrance, and skewed PCR amplification steps. All these help to preserve the fidelity of native small RNA levels and achieve the unbiased high quantification accuracy better than RNA-seq or even qPCR. Learn more>>
• Low RNA sample amount requirements
Arraystar Small RNA Microarray requires as little as 100 ng total RNA, which is magnitudes lower than what small RNA-seq requires. As its direct end labeling chemistry does not require RNA pretreatments that often cause RNA loss, the microarray significantly reduces the demand for input RNA amounts especially for heavily modified RNA biotypes (e.g. tRNA and tsRNA). The low sample amount requirement opens up opportunities for research projects where the samples are rare or of limited supply. Learn more>>
• Tolerant for RNA samples at lower qualities
The direct RNA end-labeling in Arraystar small RNA profiling is relatively insensitive to nucleotide damage in the substrate RNA sequence as it does not rely on cDNA copying by reverse transcription. This contrasts with small RNA-seq, which is vulnerable to any nucleotide damage along the small RNA sequence. Furthermore, whereas the microarray probes are unaffected by unrelated sequence presence, RNA fragments from the abundant rRNAs in degraded RNA samples can contaminate small RNAs in the size range, depressing small RNA coverage in small RNA-seq. Learn more>>
For these reasons, Small RNA Array is particularly advantageous for preserved or chemically treated samples or degraded samples. e.g. serum/plasma/biofluid/FFPE RNAs.
Human Small RNA Expression Array V1.0
Total number of distinct probes | 14,707 |
Probe design strategy | The whole probe includes 5′-cap segment, specific sequence for small |
Probe sites | miRNAs and tsRNAs: 3′-sequence of small RNAs Pre-miRNA: Loop sequence of the pre-miRNA tRNA: anti-codon loop sequence of the mature tRNA snoRNA: A specific sequence region in the snoRNA |
Probe specificity | Small RNA specific |
Coverage of miRNAs | 2,627 (1,318 5-p-miRNA and 1,309 3-p-miRNA) |
Coverage of tsRNAs | 4,254 |
Coverage of pre-miRNAs | 1,745 |
Coverage of mature-tRNAs | 346 |
Coverage of snoRNAs | 955 |
small RNA sources | miRNA: miRBase(v22) Literatures: Scientific publications up to 2019(references 1-40) |
Array Format | 8 x 15K |
Mouse Small RNA Expression Array V1.0
Total number of distinct probes | 14,192 |
Probe design strategy | The whole probe includes 5′-cap segment, specific sequence for small |
Probe sites | miRNAs and tsRNAs: 3′-sequence of small RNAs Pre-miRNA: loop sequence of the pre-miRNA tRNA: anti-codon loop sequence of the mature tRNA snoRNA: specific sequence along the entire length of the snoRNA |
Probe specificity | small RNA specific |
Coverage of miRNAs | 1,949 (966 5-p-miRNA and 983 3-p-miRNA) |
Coverage of tsRNAs | 1,767 |
Coverage of pre-miRNAs | 1,122 |
Coverage of mature-tRNAs | 270 |
Coverage of snoRNAs | 1,324 |
| small RNA sources | miRNA: miRBase(v22) Literatures: Scientific publications up to 2019(references 1-40) |
Array Format | 8 x 15K |
Rat Small RNA Expression Array V1.0
Total number of distinct probes | 14,238 |
Probe design strategy | The whole probe includes 5′-cap segment, specific sequence for small |
Probe sites | miRNAs and tsRNAs: 3′-sequence of small RNAs Pre-miRNA: loop sequence of the pre-miRNA tRNA: anti-codon loop sequence of the mature tRNA snoRNA: specific sequence along the entire length of the snoRNA |
Probe specificity | small RNA specific |
Coverage of miRNAs | 749 (355 5-p-miRNA and 394 3-p-miRNA) |
Coverage of tsRNAs | 1,135 |
Coverage of pre-miRNAs | 448 |
Coverage of mature-tRNAs | 197 |
Coverage of snoRNAs | 1,486 |
| small RNA sources | miRNA: miRBase(v22) |
Array Format | 8 x 15K |
The data analyses include profiling measurement values, statistical computations, informative annotations, and publication quality graphics.
Differential expression analysis of small RNAs
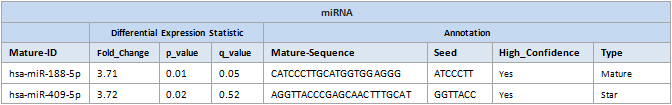

Mature-ID: miRBase ID for the mature miRNA.
FC: The fold change comparing the sample groups.
P-value: The probability value of a t-test.
q-value (FDR): The FDR adjusted p-value.
Mature-Sequence: Sequence of the mature miRNA.
Seed: Seed sequence of the mature miRNA (Positions 2-8 of the mature miRNA).
High_Confidence: A microRNA with robust support as an authentic miRNA gene.
Type: MirGeneDB annotated miRNA type based on expression level.
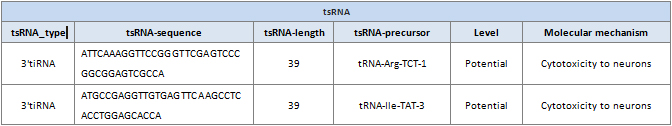
tsRNA_type: tsRNA type (tRF-5, tRF-3, tRF-1, 5-Leader, 5-tiRNA, 3-tiRNA, and i-tRF).
tsRNA-sequence: tsRNA sequence.
tsRNA-length: tsRNA length.
tsRNA-precursor: Symbol for the tsRNA precursor.
Level: Confidence level for the tsRNA
Functional – Documented with characterized biological functions or disease association;
Reliable – Recorded in tRFdb or reported by literatures, but without further studies;
Potential – Predicted by Arraystar based on RNA fragment lengths and cleavage positions in the tRNA.
Mechanism: The molecular mechanism of tsRNA.
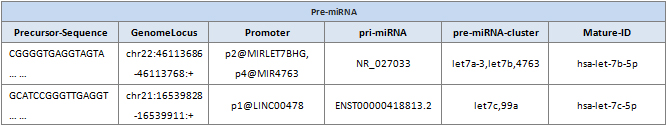
Precursor-Sequence: Sequence of the pre-miRNA.
GenomeLocus: Genome locus of the miRNA gene.
Promoter: Fantom5 promoter name of the miRNA gene.
pri-miRNA: Primary transcript of the miRNA annotated in Fantom5 (available for human and mouse only).
pre-miRNA-cluster: A group of pre-miRNAs with inter-miRNA distances <10 kb on the same chromosome and DNA strand.
Mature-ID: ID of the mature miRNA (not its star miRNA) for this pre-miRNA.
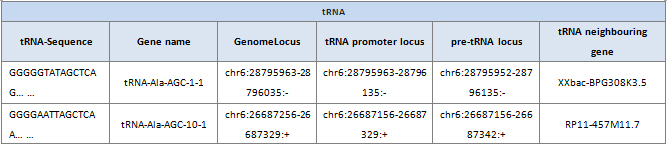
Sequence: Sequence of the tRNA isodecoder.
Gene name: Gene name of the isodecoder tRNA.
GenomeLocus: Genome locus of the tRNA isodecoder.
tRNA promoter Locus: Genome locus of the tRNA isodecoder promoter.tRNA promoter – tRNA promoters which include a tRNA gene plus 100 base pairs of upstream sequence. (PMC6108506).
pre-tRNA locus: Genome locus of the tRNA isodecoder precursor. pre-tRNA – precursor tRNA which include a tRNA gene plus 100 base pairs of upstream sequence and a 3’trailer.
tRNA neighboring gene: The nearest gene name of the tRNA isodecoder.
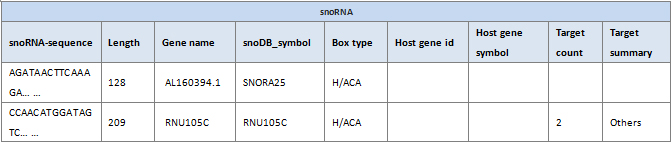
snoRNA-sequence: Sequence of the snoRNA.
Length: snoRNA length.
Gene name: Gene name of the snoRNA.
snoDB_symbol: snoDB symbol for the snoRNA (http://scottgroup.med.usherbrooke.ca/snoDB/).
Box type: snoRNA box type based on highly conserved motifs (H/ACA and C/D).
Host gene id: The ID of genes with intronic sequences overlapping that of a snoRNA.
Host gene symbol: snoRNA host gene symbol.
Target count: snoRNA target number.
Target summary: snoRNA target biotype (lncrna, mirna, ncrna, protein coding, pseudogene, rrna, snorna, snrna, trna and other).
Hierarchical clustering heatmap of differentially expressed small RNAs
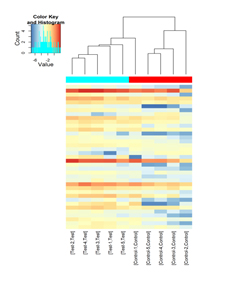
Figure 1. Hierarchical clustering heatmap of differentially expressed miRNAs. The miRNA expression levels are represented by red-blue color scale referenced in the color key on the top left. The top dendrogram shows the relative closeness of the expression profiles among the samples. The sample group membership is indicated by the color bars above the heat map.
tRF16 affects NFKBIA stability and promotes osteoarthritis progression by regulating ALKBH5 expression in m6A-dependent manner. Zhu C, et al. Communications Biology, 2025
Plasma-Derived Exosomal i-tRF-LeuCAA as Biomarker for Glioma Diagnosis and Promoter of Epithelial-Mesenchymal Transition via TPM4 Regulation. Liu H, et al. Cns Neuroscience & Therapeutics, 2025
Eurycomalactone switched hepatocellular carcinoma cells into quiescence through 5’tRFAla/DVL/ß-catenin pathway inhibition. Zhang Z, et al. Scientific Reports, 2025
tRF5–22-SerGCT-1 protects the heart against myocardial injury by targeting MSK1. Meng F, et al. Epigenomics, 2025
MicroRNA sequencing analysis in pediatric patients with influenza-associated acute necrotizing encephalopathy: Potential biomarkers for early diagnosis and therapy. Wang W, et al. Infection, Genetics and Evolution, 2025
5’tiRNA-35-GlyTCC-3 and 5’tiRNA-33-CysGCA-11 target BMP6, CUL1 and SPR of non-syndromic cleft palate. Liu R, et al. BMC Oral Health, 2025
P1.02D.02 Tumor-Derived Exosomal tsRNA 3’ tiRNA-AlaCGC Promotes Immune Tolerance by Inducing Fibroblast Senescence in LUAD. Zhang Y, et al. Journal of Thoracic Oncology, 2024
Optimizing of a suitable protocol for isolating tissue-derived extracellular vesicles and profiling small RNA patterns in hepatocellular carcinoma. Yang W,et al. Liver International, 2024
Dynamic PAH-Related Changes: A Dataset of tRNA-Derived Small RNA Transcriptome Across Multiple Organs. Chen Y, et al. Scientific Data, 2024
Serum extracellular vesicles 3’tRF-ThrCGTand 3’tRF-mtlleGAT combined with tumor markers can serve as minimally invasive diagnostic predictors for colorectal cancer. Peng J, et al. Frontiers in Oncology, 2024
Microarray analysis of tRNA-derived small RNA (tsRNA) in LPS-challenged. Lin H, et al. Gene, 2024
Small nucleolar RNA Snora73 promotes psoriasis progression by sponging miR-3074-5p and regulating PBX1 expression. Zhang L, et al. Functional & Integrative Genomics, 2024
The diagnostic value of serum exosomal SNORD116 and SNORA21 for NSCLC patients. Li L,et al. Clinical & translational oncology, 2024
Bioinformatic analysis of the expression profile and identification of RhoGDI2 as a biomarker in imati***-resistant K562 cells. Yang Y, et al. Hematology, 2023
Microarray Analysis Reveals Changes in tRNA-Derived Small RNAs (tsRNAs) Expression in Mice with Septic Cardiomyopathy. Yuan L,et al. Genes, 2022
