NGS Solutions
Related Products
Related Services
Related Reviews
RNA Modification Sequencing – m6A/m5C/m1A/ac4C/m7G/Ψ
Arraystar RNA Modification Sequencing(MeRIP-seq) profiles your choice of m6A, m1A, m5C, ac4C, m7G or Ψ epitranscriptomic modification by RNA immunoprecipitation sequencing. Please specify one modifcation per sequencing experiment when you request a quote.
For m6A-seq, we offer two m6A-RNA enrichment methods to choose from: MeRIP by m6A antibody immunoprecipitation or pull down by m6A reader GST-YTH.
Benefits:
Superb expertise: Decade of experience and skills in DNA/RNA modification profiling.
Site coverage: Detects the modification in any sites in a transcript not limited to observed motif compliance.
High resolution: Accurate peak positioning within 100 bases of the m6A sites.
High accuracy: Modification levels calibrated with the transcript abundance in the input control.
Stringent QC: High IP efficiency verified by qPCR.
Publication ready: Differential modification analysis, modification peak distribution, and motif identification.
| Service Name | Modification | Method | Price |
|---|---|---|---|
| Epitranscriptomic Sequencing Service | m6A/m5C/m1A/ac4C/m7G/Ψ *Specify one in the quote | Antibody | |
| Epitranscriptomic Sequencing Service | m6A | GST-YTH pull down |
GST-YTH is a recombinant fusion protein of YTH-DF2 m6A reader domain (a.a385-579) with a GST tag for m6A RNA enrichment. YTH is an evolutionarily conserved structural domain that selectively “reads” and binds m6A within the consensus RRACH motifs [1]. Structurally, the YTH domain contains two or three tryptophan residues that form an aromatic cage and binding pocket for m6A, with additional interactions with nucleotides before and after m6A, thus giving sequence preference to the RRACH motif [2-4](Fig.1). That is, GST-YTH binds m6A-containing RNAs in an m6A structure- and RRACH sequence motif-dependent manner [1], both of which are distinct from m6Am and other similar RNA modifications. Thus, GST-YTH pulldown is highly specific to m6A without the cross-reactivity with other structurally similar RNA modifications[5], particularly m6Am [6], as by m6A antibody MeRIP.
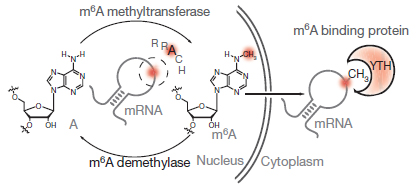
Figure 1. YTH binds to m6A-modified RNAs in an m6A structure- and RRACH sequence motif-dependent manner, at higher specificity to m6A without the cross-reactivity to other similar RNA modifications such as m6Am.
Reference
1. Wang X et al: N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 2014, 505(7481):117-120.[PMID: 24284625]
2. Luo S, Tong L: Molecular basis for the recognition of methylated adenines in RNA by the eukaryotic YTH domain. Proc Natl Acad Sci U S A 2014, 111(38):13834-13839.[PMID: 25201973]
3. Theler D et al: Solution structure of the YTH domain in complex with N6-methyladenosine RNA: a reader of methylated RNA. Nucleic Acids Res 2014, 42(22):13911-13919.[PMID: 25389274]
4. Xu C et al: Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain. Nat Chem Biol 2014, 10(11):927-929.[PMID: 25242552]
5. Linder B et al: Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods 2015, 12(8):767-772.[PMID: 26121403]
6. https://sysy.com/product/202003
RNA modification peaks in the transcript
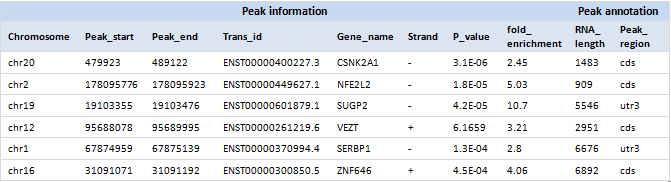
RNA modification peak visualization
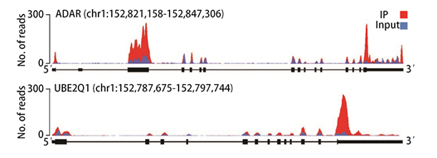
Fig 1. m6A peaks in human ADAR1 and UBE2Q1 mRNAs are visualized with the provided track files in a genome browser
Distribution of modification peak reads across mRNA regions
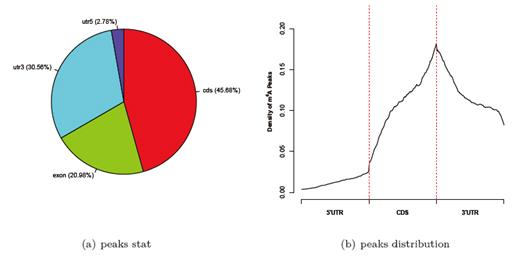
Fig 2. (a) Peak stat pie chart and (b) Peak density distribution, of modification peaks in 5’UTR, CDS, and 3’UTR regions of the mRNAs.
Motif analysis of modification peak sequences


Fig 3. Sequence motifs within the RNA modification peaks are identified with DREME algorithm and shown in sequence logo representation.
Your samples
Purified total RNA, tissues, or cells
Please refer to Sample Submission for details about how to prepare the samples and get your project started.
Arraystar Epitranscriptomic Sequencing
1. Total RNA samples (Optional service available for RNA extraction from tissues or cells)
2. RNA QC
3. mRNA isolation from the total RNA by poly(A) selection
4. mRNA fragmentation
5. Immunoprecipitation with choice of m6A, m1A, m5C, ac4C, m7G, Pseudouridine antibody
6. IP efficiency QC by qPCR
7. Sequencing library preparation
8. Cluster generation on sequencing flow cell
9. Sequencing by Illumina sequencing platform
10. Data analysis, including mapping, peak calling, peak annotation, motif analysis, differentially modifcation peak analysis, Gene Ontoloty (GO) and Pathway analyses.
YTH N6-methyladenosine RNA Binding Protein 1 Inhibits Smooth Muscle Cell Phenotypic Modulation and Neointimal Hyperplasia. Tian K, et al. cells, 2025
RNA N6-Methyladenosine Affects Copper-Induced Oxidative Stress Response in Arabidopsis thaliana. Sharma B, et al, Non-Coding RNA, 2024
m6A modification of HSATIII lncRNAs regulates temperature-dependent splicing. Ninomiya K, et al. The EMBO Journal, 2021
A defined N6-methyladenosine (m6A) profile conferred by METTL3 regulates muscle stem cell/myoblast state transitions. Gheller B J, et al. Cell Death Discovery, 2020
MTHFD2 links RNA methylation to metabolic reprogramming in renal cell carcinoma. Green N H, et al. Oncogene, 2019
