Gene Expression Array Service
Related Products
Related Services
Related Reviews
Gene Expression Array Service
Benefits:
• Our expertise in the microarray experiments and high array technical quality are the key to the success of gene expression profiling.
• Our service is based on the most powerful platform in this field, Agilent, offering variety of choices in array content, density and analysis tools.
The gene expression arrays that are most often selected by our customers include:
| Microarray | Species | Format | Probe length | Coverage |
| Whole Human Genome | Human | 4 * 44K | 60-mer | 27,958 |
| Whole Mouse Genome | Mouse | 4 * 44K | 60-mer | 39,430 |
| Whole Rat Genome | Rat | 4 * 44K | 60-mer | 26,930 |
| Service Name | Format | Price |
|---|---|---|
| Gene Expression Array Service | 4*44K | |
| Gene Expression Array Service | 8*60K |
Arraystar’s bioinformatics team has extensive experience in analyzing gene expression array data. We provide comprehensive data analyses, including:
• Raw Data Files
• Summary Report
• Box Plot, Scatter Plot and Hierarchical Clustering
• Differentially Expressed Gene Screening
• GO Analysis and Pathway Analysis(Fig.1)
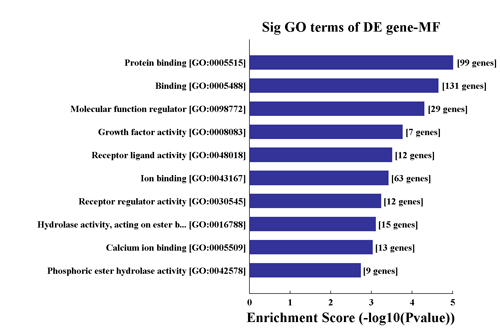
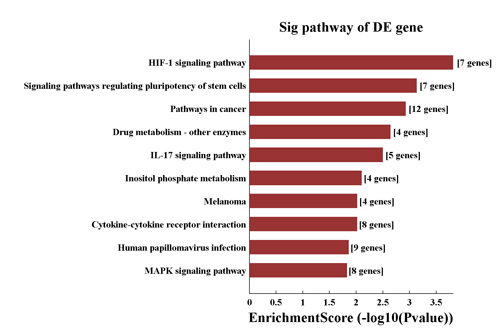
Fig.1 GO and Pathway Analysis. Significant GO terms/Pathways of differentially expressed gene. A bar plot is generated from the top 10 GO terms/Pathways containing the largest fold enrichment of differentially expressed genes. The plot gives a more intuitive view of the significant GO terms/Pathways.
Please refer to Sample Submission for details in how to get your project started.
• RNA isolation (Optional)
• RNA QC
• cDNA synthesis
• Target preparation by labeling with Cy3
• Array hybridization, washing, and scanning
• Data extraction, analysis and summarization
Identification and characterization of a novel adiponectin receptor agonist adipo anti-inflammation agonist and its anti-inflammatory effects in vitro and in vivo. Qiu W, et al. British Journal of Pharmacology, 2021
A surrogate of Roux-en-Y gastric bypass (the enterogastro anastomosis surgery) regulates multiple beta-cell pathways during resolution of diabetes in ob/ob mice. Amouyal C, et al. EBioMedicine, 2020
Dietary carbohydrates modulate metabolic and beta cell adaptation to high fat diet-induced obesity. Her T, et al. American Journal of Physiology-Endocrinology and Metabolism. 2020
Astrocytes influence medulloblastoma phenotypes and CD133 surface expression. Gronseth E, et al. PloS one, 2020
The Toxoplasma effector TEEGR promotes parasite persistence by modulating NF-?B signalling via EZH2. Braun L, et al. Nature microbiology, 2019
Application of hydroxyapatite nanoparticles in tumor-associated bone segmental defect. Zhang K, et al. Science Advances, 2019
Engineered triple inhibitory receptor resistance improves anti-tumor CAR-T cell performance via CD56. Zou F, et al. Nature communications, 2019
Lymphocyte-Derived Exosomal MicroRNAs Promote Pancreatic ß Cell Death and May Contribute to Type 1 Diabetes Development. Guay C,et al. Cell metabolism, 2018
Osteoblast-secreted factors mediate dormancy of metastatic prostate cancer in the bone via activation of the TGFßRIII-p38MAPK-pS249/T252RB pathway. Yu-Lee L Y, et al. Cancer Research, 2018
MicroRNA-Mediated Dynamic Bidirectional Shift between the Subclasses of Glioblastoma Stem-like Cells. Rooj A K, et al. Cell Reports, 2017
MicroRNA Signatures and Molecular Subtypes of Glioblastoma: The Role of Extracellular Transfer. Godlewski J, et al. Stem Cell Reports, 2017
Bmal1 is required for beta cell compensatory expansion, survival and metabolic adaptation to diet-induced obesity in mice. Rakshit K, et al. Diabetologia, 2016
FOXP3 promotes tumor growth and metastasis by activating Wnt/ß-catenin signaling pathway and EMT in non-small cell lung cancer. Yang S,et al. Molecular Cancer, 2017
Long noncoding RNA CRNDE stabilized by hnRNPUL2 accelerates cell proliferation and migration in colorectal carcinoma via activating Ras/MAPK signaling pathways. Jiang H, et al. Cell Death & Disease, 2017
In vitro expansion impaired the stemness of early passage mesenchymal stem cells for treatment of cartilage defects. Jiang T, et al. Cell Death & Disease, 2017
Circular RNAs as novel regulators of ß-cell functions in normal and disease conditions. Stoll L, et al. Molecular Metabolism, 2017
