Related Products
Related Services
Related Reviews
Seq-Star™ rRNA Removal Kit
Benefits
• High rRNA depletion efficiency
• Enrichment of both polyadenylated RNAs (e.g. mRNA) and non-polyadenylated RNAs (e.g. many long non-coding RNAs)
• rRNA removal for both intact and degraded RNA samples (e.g. FFPE RNA)
• rRNA-depleted RNAs convenient purified and collected by the magnetic beads in the kit
• Simple and fast operating procedure
| Product Name | Catalog No. | Description | Size | Price |
|---|---|---|---|---|
| Seq-Star™ rRNA Removal Kit | AS-MB-001 | Human/Mouse/Rat NGS RNA-Seq library preparation | 12 reactions | $487.00 |
Seq-Star™ rRNA Removal (H/M/R) Kit uses an enzymatic method to remove the highly abundant rRNAs in the total RNA for NGS RNA-seq library preparation. Both cytoplasmic (5S, 5.8S, 18S, 28S) and mitochondrial (12S and 16S) ribosomal RNAs from human, mouse and rat total RNA samples are efficiently depleted (Figure 1).
Compared with mRNA enrichment by oligo(dT), Seq-Star™ rRNA Removal (H/M/R) Kit selectively digests rRNAs while retaining polyadenylated mRNAs and non-polyadenylated RNA transcripts (such as many long non-coding RNAs and antisense transcripts).
Compared with rRNA removal by oligonucleotide capture on solid phase, Seq-Star™ rRNA Removal (H/M/R) Kit is capable of removing even the fragmented rRNAs in degraded total RNAs. Therefore the kit is particularly suitable for either intact or degraded RNA samples (e.g. FFPE RNA).
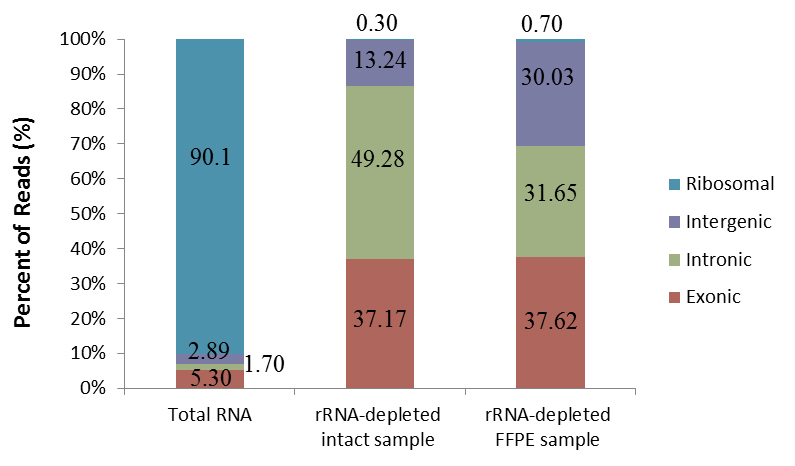
Figure1.RNA-seq results generated from intact or FFPE samples by Seq-Star™ rRNA Removal (H/M/R) Kit.
Epigenetic inhibitor zebularine activates ear pinna wound closure in the mouse. Sass P, et al. EBioMedicine, 2019
Immunophenotyping and transcriptional profiling of in vitro cultured human adipose tissue derived stem cells. Mieczkowska A, et al. Scientific reports, 2018
circRNA_100290 plays a role in oral cancer by functioning as a sponge of the miR-29 family. Chen L, et al. Oncogene, 2017
Systematic identification and comparison of expressed profiles of lncRNAs and circRNAs with associated co-expression and ceRNA networks in mouse germline stem cells. Li X, Ao J, Wu J. Oncotarget, 2017
Integrative analysis for the role of long non-coding RNAs in radiation-induced mouse thymocytes responses. Hui Gao, et al. Acta Biochimica et Biophysica Sinica, 2017

