Featured ncRNA Microarrays
Related Products
Related Services
Related Reviews
Circular RNA Array Service
Arraystar Circular RNA Arrays, systematically profiling the expression of circular RNAs, are the only commercially available tool to most effectively profile, analyze and explore this new and exciting world of noncoding RNAs. We offer the specialized Circular RNA array service, from sample to publishable data!
Benefits
• The only practical platform for highly sensitive and specific circular RNA profiling. Learn more>>
• Circular junction-specific probes, linear RNA removal and efficient circRNA labeling ensure the most specific, accurate and reliable circRNA expression profiling.
• Detailed annotation specific to circRNA biology, e.g. miRNA binding sites, miRSVR scores and conservation status, to unravel functional roles as miRNA sponges.
• The preferred choice over RNA-sequencing, due to the particular properties of circular RNA. Learn more>
Watch Video> The Latest Highlights on CircRNA in Cancer
| Service Name | Catalog No | Description | Format | Price |
|---|---|---|---|---|
| Human CircRNA Array Service | AS-S-CR-H-V2.0 | 13,617 CircRNAs | 8*15K | |
| Mouse CircRNA Array Service | AS-S-CR-M-V2.0 | 14,236 CircRNAs | 8*15K | |
| Rat CircRNA Array Service | AS-S-CR-R-V2.0 | 14,145 CircRNAs | 8*15K |
• Specific Circular Junction Probes
Reliably and accurately identify individual circRNAs, even in the presence of their linear counterparts (Fig. 1).
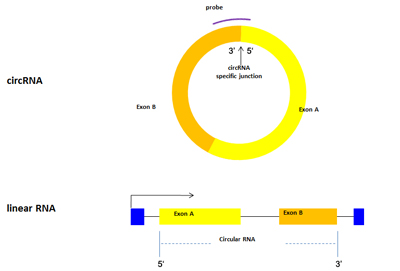
Figure 1. Arraystar circRNA Array V2.0 uses specific circular junction probes to accurately and reliably detect each individual circRNAs, even in the presence of their linear counterparts. The linear RNA at the bottom is alternatively processed to generate a circular variant above. A probe is designed to target the circRNA-specific junction site, where the 5′ end of exon A joins together with the 3′ end of exon B.
• Detailed annotation for circRNA-miRNA association
Annotation of potential miRNA target sites on the circular RNAs helps to unravel their functional roles as a natural miRNA sponge (Fig. 2).
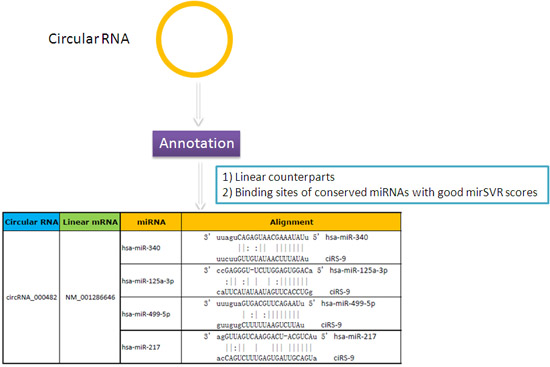
Figure 2. The association between circular RNA and conserved miRNAs is annotated in detail.
• The preferred choice over RNA-sequencing, as RNA-seq is currently inadequate for such task due to the particular properties of circular RNA. Learn more >
1) Circular junction read counts are only a fraction of the circular RNA. They are much lower than the linear RNA at the same abundance level.
2) For detection of the circular RNA presence, a few reproducible counts are required. But for quantification, at least hundreds of read counts are required.
3) Even with combined large datasets and studies, most of the known circular RNAs have just a few read counts. Count numbers at these levels are not nearly sufficient for differential expression analysis.
In short, circular RNA sequencing can be used for novel discovery by a few read counts, but is inadequate for differential expression analysis even at modest abundance levels.
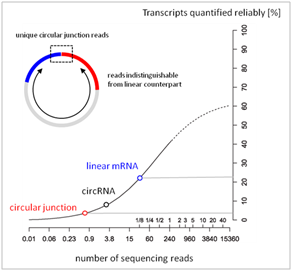
Figure 3. RNA-seq quantification reliability vs read depth. Typical RNA-seq has a depth of < 30 mil reads for mRNAs (blue circle), which is < 0.5 mil for cross circular junction reads (red circle). Less than 5% circular junctions can be reliably quantified. Adopted from Labaj et al, (2011) Bioinformatics [PMID 21685096].
• Guaranteed performance
– Better sensitivity: Low abundance RNAs are accurately detected with a wide dynamic range of over 5 orders of magnitude (Fig. 4).
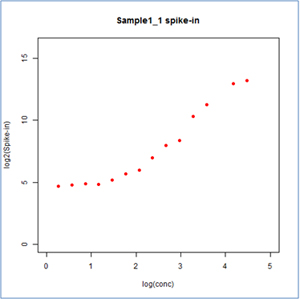
Figure 4. A wide dynamic detection range of over 5 orders of magnitude with Arraystar’s circRNA Arrays.
– High Reproducibility: Technical replicates show tight correlation on Arraystar circRNA Arrays (R2>0.9) (Fig. 5).
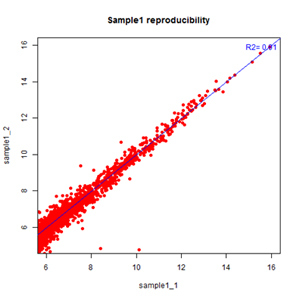
Figure 5. High reproducibility on Arraystar circRNA arrays.
Circular RNA (circRNA) is a novel type of RNA that, unlike linear RNA, forms a covalently closed continuous loop, some of which are highly represented in the eukaryotic transcriptome. Most of these circRNAs are generated from exonic or intronic sequences, are conserved across species, and often show tissue/developmental-stage-specific expression. Circular RNAs are more stable than linear RNAs owing to their higher nuclease stability, which constitutes an enormous advantage from a clinical point of view as a novel class of biomarkers. In addition, circRNAs have been shown to function as natural miRNA sponge transcripts, the so-called competing endogenous RNAs (ceRNAs) in diverse species. Their interaction with disease associated miRNAs suggests the potential importance of circular RNAs in disease regulation.
To facilitate the analysis of circRNAs, Arraystar has developed the first commercially-available circRNA arrays for human , mouse and rat. In order to detect circRNAs comprehensively and reliably, we have updated the circRNA repertoire represented in the previous Version 1.0 and launched the newly designed V2.0.
It is the only practical platform for highly sensitive and specific circular RNA profiling available for biologists with a complete from RNA sample to publishable data service. To learn more about the publications by our clients, click here.
Human Database
| Total Number of Distinct Probes | 13,617 |
| Probe Length | 60 nt |
| Probe Selection Region | Probes targeting circRNA-specific junctions |
| Probe Specificity | Transcript-specific |
| Labeling Method | Random primer labeling coupled with RNase R sample pretreatment to ensure specific and efficient labeling of circular RNAs. |
| Circular RNA Sources | |
| Salzman’s circRNAs [4] | 8,529 |
| Memczak’s circRNAs [3] | 1,601 |
| Zhang’s circRNAs [6] | 93 |
| Zhang’s circRNAs [5] | 4,980 |
| Jeck’s circRNAs [2] | 3,769 |
| Guo’s circRNAs [1] | 5,536 |
| Array Format | 8 * 15K |
Mouse Database
| Total Number of Distinct Probes | 14,236 |
| Probe Length | 60 nt |
| Probe Selection Region | Probes targeting circRNA-specific junctions |
| Probe Specificity | Transcript-specific |
| Labeling Method | Random primer labeling coupled with RNase R sample pretreatment to ensure specific and efficient labeling of circular RNAs. |
| CircRNA Sources | |
| Memczak’s circRNAs | 1,750 |
| Guo’s circRNAs | 570 |
| You’s circRNAs | 13,300 |
| Array Format | 8 * 15K |
Rat Database
| Total Number of Distinct Probes | 14,145 |
| Probe Length | 60 nt |
| Probe Selection Region | Probes targeting circRNA-specific junctions |
| Probe Specificity | Transcript-specific |
| Labeling Method | Random primer labeling coupled with RNase R sample pretreatment to ensure specific and efficient labeling of circular RNAs. |
| CircRNA Sources | |
| You Xintian’s circRNAs [7] | 12,298 |
| Mouse circRNA orthologs | 1,668 |
| Human circRNA orthologs | 179 |
| Array Format | 8 * 15K |
References
1. Guo, J. U., V. Agarwal, et al. (2014). “Expanded identification and characterization of mammalian circular RNAs.” Genome Biol 15(7): 409.
2. Jeck, W. R., J. A. Sorrentino, et al. (2013). “Circular RNAs are abundant, conserved, and associated with ALU repeats.” RNA 19(2): 141-157.
3. Memczak, S., M. Jens, et al. (2013). “Circular RNAs are a large class of animal RNAs with regulatory potency.” Nature 495(7441): 333-338.
4. Salzman, J., R. E. Chen, et al. (2013). “Cell-type specific features of circular RNA expression.” PLoS Genet 9(9): e1003777.
5. Zhang, X. O., H. B. Wang, et al. (2014). “Complementary sequence-mediated exon circularization.” Cell 159(1): 134-147.
6. Zhang, Y., X. O. Zhang, et al. (2013). “Circular intronic long noncoding RNAs.” Mol Cell 51(6): 792-806.
7. You X., I. Vlatkovic, et al. (2015). “Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity.” Nat Neurosci 18(4): 603-610.
Please refer to Sample Submission for details in how to get your project started.
• RNA isolation (optional)
• RNA QC
• Determine the purity and concentration of total RNA
• Assess the integrity of total RNA
• RNase R treatment
• cDNA synthesis
• Labeling
• Array hybridization, washing, and scanning
• Data extraction, analysis and summarization
• Streamlined and optimized labeling pipeline
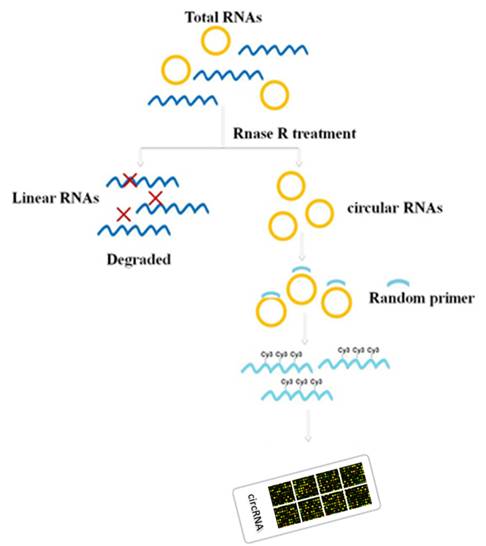
Figure 1. A random primer-based labeling system is coupled with RNase R-based sample pretreatment to efficiently remove linear RNAs, and specifically label circular RNAs.
• Detailed annotation for circRNA-miRNA association
Annotates the circRNAs with the potential target sites of miRNAs, which will be helpful for unraveling their functional roles as a natural miRNA sponge.
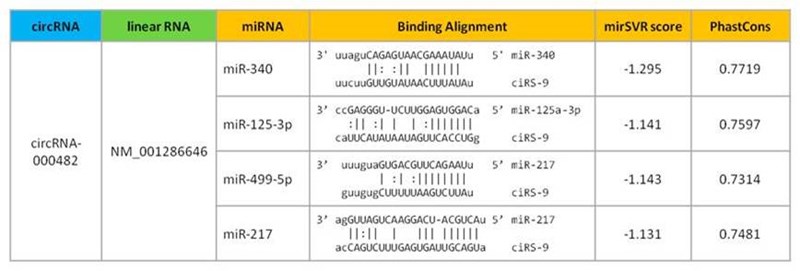
Figure 1. The circular RNA and its asscociated conserved miRNAs are annotated in detail.
• Differentially Expressed circRNAs
Differentially expression circRNAs are selected by the magnitude of change and the statistic significance in p-value, which are visualized on Volcano plots. The expression patterns are displayed on hierarchical clustering heatmaps.
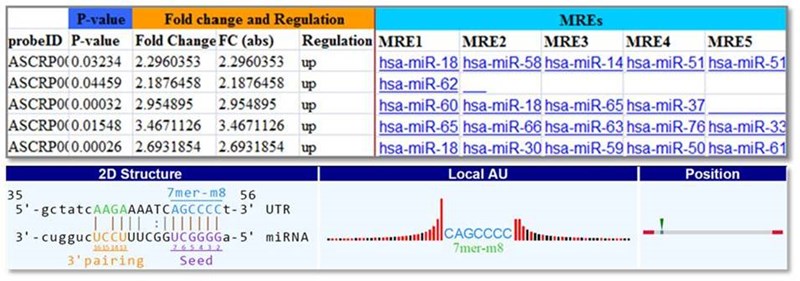

Figure 2. Differentially expressed circRNAs, the binding with multiple microRNAs, and the details of the microRNA Response Elements (MREs).
Microglial circDlg1 modulates neuroinflammation by blocking PDE4B ubiquitination-dependent degradation associated with Alzheimer’s disease. Shi J, et al. Theranostics, 2025
Atrial remodelling and atrial fibrillation self-sustaining: the role of circulating circDGCR8. Gao Y, et al. Cardiovascular Research, 2025
Circular RNA circPHLPP2 promotes tumor growth and anti-PD-1 resistance through binding ILF3 to regulate IL36 gamma transcription in colorectal cancer. Hu Y, et al. Molecular Cancer, 2024
CircPDIA3/miR-449a/XBP1 feedback loop curbs pyroptosis by inhibiting palmitoylation of the GSDME-C domain to induce chemoresistance of colorectal cancerRunning Head: circRNA curbs pyroptosis to induce chemoresistance. Lin J,et al. Drug Resistance Updates, 2024
Droplets Cas13a-RPA measurement delineates potential role for plasma circWDR37 in colorectal cancer. Xu J, et al. Aggregate, 2024
circMIRIAF aggravates myocardial ischemia-reperfusion injury via targeting miR-544/WDR12 axis. Yin L, et al. Redox biology, 2024
circFAM193B interaction with PRMT6 regulates AML leukemia stem cells chemoresistance through altering the oxidative metabolism and lipid peroxidation. Yang X, et al. Leukemia, 2024
CircPDE5A-encoded novel regulator of the PI3K/AKT pathway inhibits esophageal squamous cell carcinoma progression by promoting USP14 mediated deubiquitination of PIK3IP1. Lei K, et al. Journal of Experimental & Clinical Cancer Research, 2024
A tumor suppressor protein encoded by circKEAP1 inhibits osteosarcoma cell stemness and metastasis by promoting vimentin proteasome degradation and activating anti-tumor immunity. Zhang Y, et al, Journal of Experimental & Clinical Cancer Research, 2024
CircPTEN suppresses human clear cell renal carcinoma progression and resistance to mTOR inhibitors by targeting epigenetic modification. Zhan Y, et al. Drug Resistance Updates, 2023
TGF-ß signaling promotes cervical cancer metastasis via CDR1as. Zhong G, et al. Molecular Cancer, 2023
CircRNA DICAR as a novel endogenous regulator for diabetic cardiomyopathy and diabetic pyroptosis of cardiomyocytes. Yuan Q, et al. Signal Transduction and Targeted Therapy, 2023
CircNOLC1 Promotes Colorectal Cancer Liver Metastasis by Interacting with AZGP1 and Sponging miR-212-5p to Regulate Reprogramming of the Oxidative Pentose Phosphate Pathway. Yuan M, et al. Advanced Science, 2023
An interaction between eIF4A3 and eIF3g drives the internal initiation of translation. Hang J, et al. Nucleic Acids Research, 2023
CircOGDH Is a Penumbra Biomarker and Therapeutic Target in Acute Ischemic Stroke. Liu Y, et al. Circulation Research, 2022
CircGPR137B/miR-4739/FTO feedback loop suppresses tumorigenesis and metastasis of hepatocellular carcinoma. Liu L, et al. Molecular Cancer, 2022
The N6-methyladenosine modification of circALG1 promotes the metastasis of colorectal cancer mediated by the miR-342-5p/PGF signalling pathway. Lin C, et al. Molecular Cancer, 2022
Hsa_circ_0003258 promotes prostate cancer metastasis by complexing with IGF2BP3 and sponging miR-653-5p. Yu Y Z, et al. Molecular Cancer, 2022
circCAPRIN1 interacts with STAT2 to promote tumor progression and lipid synthesis via upregulating ACC1 expression in colorectal cancer. Yang Y, et al. Cancer Communications, 2022
The circular RNA circDLG1 promotes gastric cancer progression and anti-PD-1 resistance through the regulation of CXCL12 by sponging miR-141-3p. Chen D L, et al. Molecular Cancer, 2021
Circular RNA circIPO11 drives self-renewal of liver cancer initiating cells via Hedgehog signaling. Gu Y, et al. Molecular Cancer, 2021
CircRNF220, not its linear cognate gene RNF220, regulates cell growth and is associated with relapse in pediatric acute myeloid leukemia. Liu X, et al. Molecular Cancer, 2021
The circACTN4 interacts with FUBP1 to promote tumorigenesis and progression of breast cancer by regulating the expression of proto-oncogene MYC. Wang X, et al. Molecular Cancer, 2021
CircMEMO1 modulates the promoter methylation and expression of TCF21 to regulate hepatocellular carcinoma progression and sor*****b treatment sensitivity. Dong Z R, et al. Molecular Cancer, 2021
Circular RNA ACTN4 promotes intrahepatic cholangiocarcinoma progression by recruiting YBX1 to initiate FZD7 transcription. Chen Q, et al. Journal of Hepatology, 2021
Circular RNA circHIPK3 promotes homeostasis of the intestinal epithelium by reducing miR-29b function. Xiao L, et al. Gastroenterology, 2021
Targeting Mitochondria-Located circRNA SCAR Alleviates NASH via Reducing mROS Output. Qiyi Zhao, et al. Cell, 2020
Extracellular Vesicle-Mediated Delivery of CircSCMH1 Promotes Functional Recovery in Rodent and Nonhuman Primate Ischemic Stroke Models. Yang L, et al. Circulation, 2020
1972P Circular RNA is associated with enz***amide resistant prostate cancer. Lim M C J, et al. Annals of Oncology, 2020
Circular RNAs as biomarkers in liquid biopsy in colorectal cancer. Valladares-Ayerbes M, et al. Journal of Clinical Oncology, 2020
A psychiatric disease-related circular RNA controls synaptic gene expression and cognition. Zimmerman A J, et al. Molecular Psychiatry, 2020
CircPTK2 (hsa_circ_0005273) as a novel therapeutic target for metastatic colorectal cancer. Yang H, et al. Molecular Cancer, 2020
Circ-AKT3 inhibits clear cell renal cell carcinoma metastasis via altering miR-296-3p/E-cadherin signals. Xue D, et al. Molecular Cancer, 2019
circGSK3ß promotes metastasis in esophageal squamous cell carcinoma by augmenting ß-catenin signaling. Hu X, et al. Molecular Cancer, 2019
A Noncoding Regulatory RNAs Network Driven by Circ-CDYL Acts Specifically in the Early Stages Hepatocellular Carcinoma. Wei Y, et al. Hepatology, 2019
Circular RNA CircFndc3b modulates cardiac repair after myocardial infarction via FUS/VEGF-A axis. Garikipati V., et al. Nature Communications, 2019
FOXP1 circular RNA sustains mesenchymal stem cell identity via microRNA inhibition. Cherubini A, et al. Nucleic Acids Research, 2019
Circular RNA cESRP1 sensitises small cell lung cancer cells to chemotherapy by sponging miR-93-5p to inhibit TGF-ß signalling. Huang W, et al. Cell Death & Differentiation, 2019
A Circular RNA Protects Dormant Hematopoietic Stem Cells from DNA Sensor cGAS-Mediated Exhaustion. Xia P, et al. Immunity, 2018
PRMT5 circular RNA promotes metastasis of urothelial carcinoma of the bladder through sponging miR-30c to induce epithelial-mesenchymal transition. Chen X, et al. Clinical Cancer Research, 2018
circEPSTI1 as a prognostic marker and mediator of triple-negative breast cancer progression. Chen B, et al. Theranostics, 2018
Circular RNA MTO1 acts as the sponge of miR-9 to suppress hepatocellular carcinoma progression. Han D, et al. Hepatology (Baltimore, Md.), 2017
A Circular RNA Binds To and Activates AKT Phosphorylation and Nuclear Localization Reducing Apoptosis and Enhancing Cardiac Repair. Zeng Y, et al. Theranostics, 2017
circRNA Mediates Silica-Induced Macrophage Activation Via HECTD1/ZC3H12A-Dependent Ubiquitination. Zhou Z, et al. Theranostics, 2017
