Small RNA NGS Solutions
Related Products
Related Services
Related Reviews
microRNA Sequencing Service
Arraystar offers a complete microRNA-seq service from RNA preparation and QC, library construction, sequencing, to comprehensive data analyses.
Benefits
Technological Performance
• Highly optimized procedures to maximize the mappable reads and deliver precise quantification
• Identification of isomiRs, and distinction of mature miRNAs from pre-microRNAs
• No cross-hybridization ambiguity issues
• Sensitive detection for low abundance miRNAs at single-copy per cell level
• Cost-effective highly optimized full service from Sample to Data
Coverage and Annotation
• Applicable to any species, not dependent on prior miRNAs reference
• Always annotated with the latest miRBase
Advanced Analyses
• Novel miRNA discovery and secondary structure prediction
• Target gene prediction and network
• Gene ontology/pathway analysis of miRNA Target Genes
| Service Name | Price |
|---|---|
| microRNA Sequencing Service |
Novel miRNA discovery and secondary structure prediction (Fig. 1)
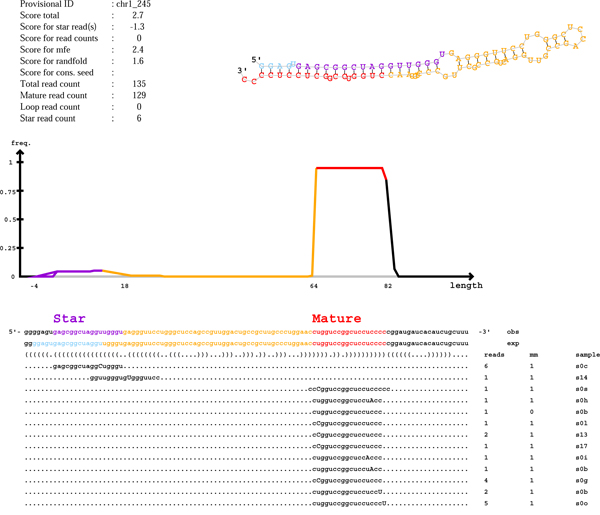
Target gene prediction and network (Fig. 2)
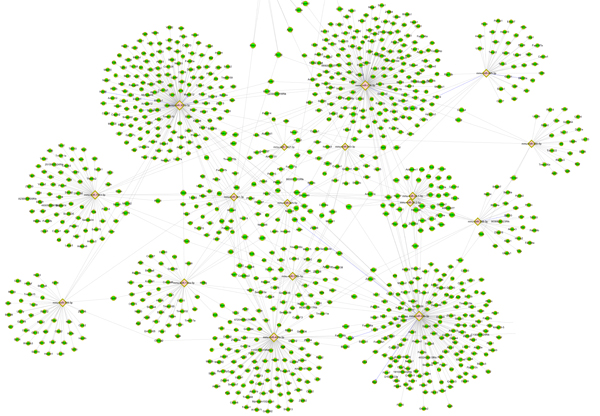
Gene ontology/pathway analysis of miRNA Target Genes (Fig. 3)
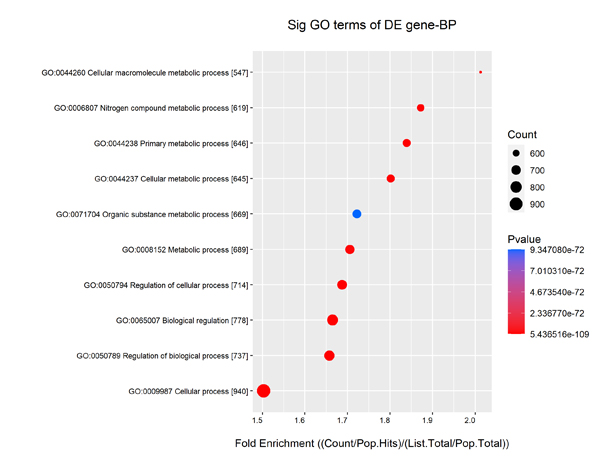
Your samples
Purified total RNA, or frozen tissues, or cell pellets
Please refer to Sample Submission for details in how to get your project started.
Arraystar microRNA seq
1. RNA isolation (Optional)
2. RNA QC
3. Sequencing library construction and library QC
4. Sequencing by illumina platform
5. Raw read data processing
6. Data analyses and report
7. Project data delivery
Cellular hierarchies of embryonal tumors with multilayered rosettes are shaped by oncogenic microRNAs and receptor–ligand interactions. Alexander Beck, et al. Nature Cancer, 2025
Irradiation alters extracellular vesicle microRNA load in the serum of patients with leukaemia. Stephanie Hehlgans, et al. Strahlentherapie und Onkologie, 2024
1a,25-Dihydroxyvitamin D3 Regulates microRNA Packaging in Extracellular Matrix Vesicles and Their Release in the Matrix. Asmussen N C,et al. Calcified Tissue International, 2023
Changes in subcutaneous adipose tissue microRNA expression in response to exercise training in obese African women. Pheiffer C,et al. Scientific Reports, 2022
Re-expression of miR-200s in claudin-low mammary tumor cells alters cell shape and reduces proliferation and invasion potentially through modulating other miRNAs and SUZ12 regulated genes. Simpson K, et al. Cancer Cell International, 2021
Extracellular vesicles derived from myocardial infarction plasma inhibit BMSCs apoptosis and enhance cardiac function via AKT signaling pathway. Jin P, et al. International Immunopharmacology, 2021
Two novel microRNAs and their association with absolute blood pressure parameters in an urban South African community. Matshazi D M, et al. Molecular Biology Reports, 2021
Complexity of the microRNA transcriptome of cow milk and milk-derived extracellular vesicles isolated via differential ultracentrifugation. Benmoussa A, et al. Journal of Dairy Science, 2019
Neuronal microRNA regulation in Experimental Autoimmune Encephalomyelitis. Juzwik C A, et al. Scientific reports, 2018
Defining the rhesus macaque placental miRNAome: Conservation of expression of placental miRNA clusters between the macaque and human. Schmidt J K, et al. Placenta, 2018
MicroRNA Expression Varies according to Glucose Tolerance, Measurement Platform, and Biological Source. Dias S,et al. BioMed Research International, 2017
Next-generation sequencing of the basal cell carcinoma miRNome and a description of novel microRNA candidates under neoadjuvant vis******* therapy: an integrative molecular and surgical case study.Sand M, et al.Ann Oncol,2016
Integrated analysis of dysregulated ncRNA and mRNA expression profiles in humans exposed to carbon nanotubes. Shvedova A A, et al. PloS one, 2016
Survival and Diversity of Human Homologous Dietary MicroRNAs in Conventionally Cooked Top Sirloin and Dried Bovine Tissue Extracts. Dever J T, et al. PloS one, 2015
RNA sequencing identifies specific PIWI-interacting small non-coding RNA expression patterns in breast cancer. Hashim A,et al. Oncotarget, 2014
Molecular fingerprinting of the podocyte reveals novel gene and protein regulatory networks. Melanie Boerries, et al. Kidney International, 2013
