NGS Solutions
Related Products
Related Services
Related Reviews
RNA Sequencing Service
Arraystar integrated RNA Sequencing for mRNA provides the full service from RNA samples, library construction, sequencing runs, to comprehensive data analysis.
Benefits
• Perfected and optimized sample prep, library construction and RNA-seq procedures.
• High efficiency, high quality, cost-effective, unbiased library construction.
• dUTP chemistry to ensure extreme transcript strand-specificity.
• Ultra-high sequencing data quality to maximize the mappable reads.
• Standard analysis package includes advanced in-depth analyses that go beyond the “standard”: Novel gene/transcript discovery, Gene Set Enrichment Analysis (GSEA) for functional prediction, alternative splicing events, and more.
For high integrity intact RNA samples, the service includes poly(A) selection to purify the mRNA in the standard procedure.
For degraded RNA samples, the alternative rRNA depletion procedure to remove the highly abundant rRNAs from sequencing is required separately.
*For expression profiling of human/mouse/rat lncRNAs, we recommend specialized Arraystar LncRNA Array(H/M/R) at higher sensitivity, accuracy, and lncRNA analysis power.
| Service Name | Data Qty | Read Length | Seq Depth | Application | Price |
|---|---|---|---|---|---|
| RNA Sequencing (polyA selection) | 6G | 150bp Paired -end | 40M (20M fragments) | Excellent coverage for protein coding mRNAs | |
| RNA Sequencing (polyA selection) | 12G | 150bp Paired -end | 80M (40M fragments) | Best for alternatively spliced/low abundance mRNAs | |
| RNA Sequencing (rRNA removal) | 12G | 150bp Paired -end | 80M (40M fragments) | Covers mRNAs&lncRNAs for species other than H/M/R* |
All RNA-seq projects include:
• Raw read data with read quality filtering.
• Adapter trimming.
• Sequencing QC stats.
RNA-seq with Analysis Package includes:
(Available for human, mouse, rat, or species with established genome/transcriptome)
• Expression profiling and differential analysis at gene and transcript levels
• Correlation Matrix, Principal Component Analysis (PCA) and hierarchical clustering heatmaps to visualize sample expression correlation, distances and clusters within and between the groups
• Gene ontology and pathway analyses to explore the differentially expressed genes participating in particular biological functions or pathways.
• Gene Set Enrichment Analysis (GSEA) analyzes the genes in functional sets that tend to show expressional changes, even if their individual differential expression may not appear strong or significant.
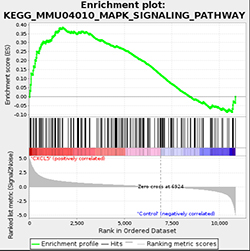
Fig 1. GSEA Enrichment Plot of MAPK signaling pathway.
• Alternative Splicing Event Detection identifies and discovers novel splice isoforms using junction sequence information with the pair-end reads and deep coverage. Different types of splicing events such as exon skipping, alternative splice sites and retained introns are detected and profiled for altered RNA processing.
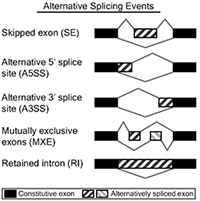
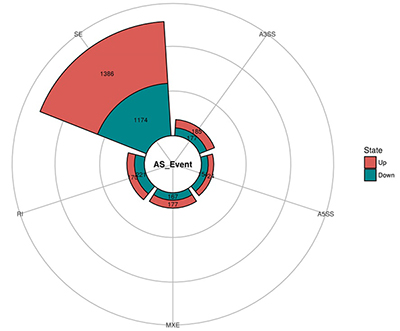
Fig 2. Alternative Splicing events classification. Fig 3. Alternative Splicing summary plot.
Your samples
Purified total RNA, or Frozen tissues, or cell pellets
Please refer to Sample Submission for details in how to get your project started.
Arraystar RNA-seq
1. Total RNA isolation (Optional)
2. RNA QC
3. mRNA enrichment by oligo-dT, or separate rRNA depletion for degraded RNA/lncRNA samples
4. Sequencing library construction and library QC
5. Sequencing by illumina platform
6. Raw read data processing
7. Data analyses and report
8. Project data delivery
Circulating microRNA analysis in a prospective co-clinical trial identifies MIR652-3p as a response biomarker and driver of regorafe*** resistance mechanisms in colorectal cancer. Hedayat S, et al. Clinical Cancer Research, 2024
BIGH3 mediates apoptosis and gap junction failure in osteocytes during renal cell carcinoma bone metastasis progression. Pan T, et al. Cancer Letters?, 2024
The hepatokine orosomucoid 2 mediates beneficial metabolic effects of bile acids. Lee S H, et al. Diabetes, 2024
Novel Effects of Prohibitin 1 Expression Level on Cholesterol and Lipid Homeostasis. Jung S, et al. The Journal of Nutritional Biochemistry, 2024
Targeting stromal cell sialylation reverses T cell-mediated immunosuppression in the tumor microenvironment. Egan H, et al. Cell Reports, 2023
Single cell G-protein coupled receptor profiling of Transcription factor 21 expressing activated kidney fibroblasts. Kaur H, et al. British Journal of Pharmacology, 2023
ING1 inhibits Twist1 expression to block EMT and is antagonized by the HDAC inhibitor vorinostat. Yang Y, et al. European Journal of Cell Biology, 2023
Decreased CRISPLD2 expression impairs osteogenic differentiation of human mesenchymal stem cells during in vitro expansion. Rong W, et al. Journal of Cellular Physiology, 2023
In vivo partial reprogramming by bacteria promotes adult liver organ growth without fibrosis and tumorigenesis. Hess S,et al. Cell Reports Medicine, 2022
Retinoic acid receptor activation reduces metastatic prostate cancer bone lesions by blocking the endothelial-to-osteoblast transition. Yu G, et al. Cancer Research, 2022
Reduction of lamin B receptor levels by miR-340-5p disrupts chromatin, promotes cell senescence and enhances senolysis. Herman A B, et al. Nucleic Acids Research, 2021
Statins reduce castration-induced bone marrow adiposity and prostate cancer progression in bone. Pan T, et al. Oncogene, 2021
FGL2-wired macrophages secrete CXCL7 to regulate the stem-like functionality of glioma cells. Yan J,et al. Cancer Letters, 2021
Molecular pathology associated with altered synaptic transcriptome in the dorsolateral prefrontal cortex of depressed subjects. Yoshino Y, et al. Translational Psychiatry, 2021
De novo transcriptome assembly of the Southern Ocean copepod Rhincalanus gigas sheds light on developmental changes in gene expression. Berger C A, et al. Marine Genomics, 2021
Cellular sources of IL-6 and associations with clinical phenotypes and outcomes in PAH. Simpson C E, et al. European Respiratory Journal, 2020
Ins****-like growth factor binding protein-2: a new circulating indicator of pulmonary arterial hypertension severity and survival. Yang J, et al. BMC medicine, 2020
The Role of Crosstalk between AR3 and E2F1 in Drug Resistance in Prostate Cancer Cells. Xu J, et al. Cells, 2020
RNA-seq Analysis of Wild-Type vs. FOXC2-Deficient Melanoma Cells Reveals a Role for the FOXC2 Transcription Factor in the Regulation of Multiple Oncogenic Pathways. Hargadon K M, et al. Frontiers in Oncology, 2020
Profiling the microRNA signature of the peripheral sensory ganglia in experimental autoimmune encephalomyelitis (EAE). Friedman T N, et al. Journal of neuroinflammation, 2019
MYC deregulates TET1 and TET2 expression to control global DNA (hydroxy) methylation and gene expression to maintain a neoplastic phenotype in T-ALL. Poole C J, et al. Epigenetics & chromatin, 2019
Subendothelial stiffness alters endothelial cell traction force generation while exerting a minimal effect on the transcriptome. Bastounis E E, et al. Scientific Reports, 2019
Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Vlachogiannis G, et al. Science, 2018
