Small RNA NGS Solutions
Related Products
Related Services
Related Reviews
tRF&tiRNA Sequencing Service
With the expertise in tRF&tiRNA, Arraystar tRFs&tiRNA sequencing service offers:
• Performance optimized tRFs&tiRNAs sequencing method with Arraystar rtStar™ tRF&tiRNA Pretreatment Kit (Arraystar, Cat.No. AS-FS-005)
• Precise annotation and classification system based on tRNA topology and statistical significance.
• Comprehensive tRF & tiRNA collection from all databases.
• tRFs&tiRNAs focused bioinformatics and statistics analyses
• Additional miRNA/piRNA*/sdRNA* expression and differential analyses (* For human, mouse, and rat species only)
• Data visualization with publication quality graphics
• Work well together with our tRNA series of technologies: tRF&tiRNA-seq, tRNA-seq, tRF&tiRNA PCR array, and tRNA PCR array
| Service Name | Price |
|---|---|
| tRF&tiRNA Sequencing Service |
tRFs and tiRNAs, generated through precise biogenesis processes from tRNA, perform many biological functions as small noncoding RNAs and are associated with many diseases and conditions (Fig.1) . They are known to act as microRNAs in RNA interference; bind protein factors to regulate target mRNA stability; interact with cytochrome c to modulate apoptosis; alter embryonic transcriptional cascades as paternal epigenetic factor in intergenerational inheritance of metabolic disorders; and assemble stress granules in response to stress conditions.
The composition and abundance of tRFs&tiRNAs are highly dependent on the cell type and disease condition, making them excellent biomolecules for biomarkers [1]. For example, the ratio of tRFs&tiRNAs is a good indicator of cancer progression-free survival and a candidate prognostic marker [2]. Also tRNA and tRF&tiRNA populations are highly enriched in biofluids, much more so than microRNAs [3-4].
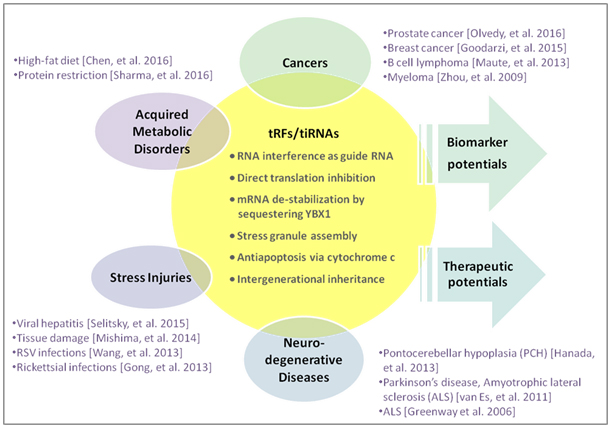
Figure 1. tRF&tiRNA functions and association with diseases.
Arraystar tRF&tiRNA sequencing service offers the sample-to-data solution. With the wealth of information from our tRF&tiRNA-seq profiling, investigators can easily advance their next step research in this new field of study, following the approaches and applications in the Roadmap for tRF &tiRNA studies.
References
1.Telonis A.G. et al. (2015) “Dissecting tRNA-derived fragment complexities using personalized transcriptomes reveals novel fragment classes and unexpected dependencies.” Oncotarget 6(28):24797-822 [PMID: 26325506]
2.Olvedy M. et al. (2016) “A comprehensive repertoire of tRNA-derived fragments in prostate cancer.” Oncotarget [PMID: 27015120]
3.Schageman J. et al. (2013) “The complete exosome workflow solution: from isolation to characterization of RNA cargo.” Biomed Res Int 2013:253957 [PMID: 24205503]
4.Dhahbi J.M. et al. (2013) “5′ tRNA halves are present as abundant complexes in serum, concentrated in blood cells, and modulated by aging and calorie restriction.” BMC Genomics 14:298 [PMID: 23638709]
The tRF&tiRNA-seq provides a wealth of bioinformatics analyses to better understand their biology and facilitate biomarker applications. As a bonus, microRNA expression analysis is also included.
Differential expression profiling and detailed annotation
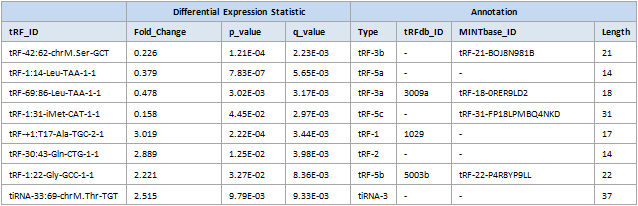
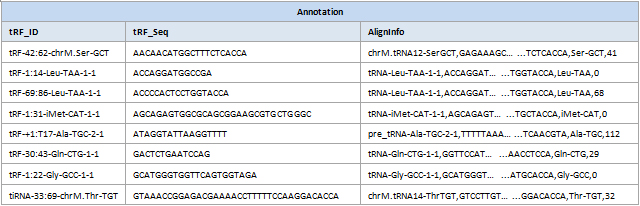
Figure 1. Differential analysis data tables with detailed annotation of tRF&tiRNA sequences, type, length and cleavage sites in tRNAs.
Pie Chart of tRF & tiRNA Subtypes
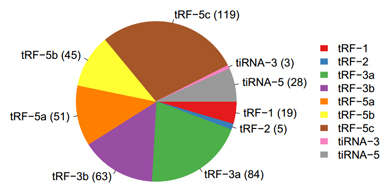
Figure 2. The distribution of tRF & tiRNA subtypes. The colors represent the tRF & tiRNA subtypes. The values in brackets represent the numbers of tRF & tiRNA subtypes.
Small RNAs derived from tRNA fragmentation regulate the functional maturation of neonatal ß cells. Bayazit M B, et al. Cell Reports, 2022
Neuronal Nsun2 deficiency produces tRNA epitranscriptomic alterations and proteomic shifts impacting synaptic signaling and behavior. Blaze J, et al. Nature Communications, 2021
Structure of human cytomegalovirus virion reveals host tRNA binding to capsid-associated tegument protein pp150. Liu Y T, et al. Nature Communications, 2021
m5U54 tRNA Hypomodification by lack of TRMT2A Drives the Generation of tRNA-Derived Small RNAs. Pereira M, et al. International Journal of Molecular Sciences, 2021
Processing by RNase 1 forms tRNA halves and distinct Y RNA fragments in the extracellular environment. Nechooshtan G,et al. Nucleic Acids Research, 2020
