Small RNA NGS Solutions
Related Products
Related Services
Related Reviews
tRNA Sequencing Service
Arraystar tRNA sequencing offers sample-to-data service to accurately profile tRNAs and sensitively detect tRNA mutations/variants/isoforms. Complete with sequencing data processing and a wealth of tRNA bioinformatic analysis, the first and only commercially available service will spearhead your tRNA research in canonical and non-canonical tRNA functions and diseases.
Benefits
• Efficiently remove internal and terminal modifications in the tRNA to reach exceptional level of tRNA-seq efficiency and accuracy using Arraystar rtStar™ tRNA Pretreatment Kit
• Performance optimized tRNA sequencing methodologies with rigorous QC process
• Comprehensive tRNA transcriptome reference from all authoritative databases
• Thorough tRNA annotation and bioinformatic analysis
• Data visualization with publication quality graphics
• Work well together with our broad portfolio of tRNA research technologies: tRF&tiRNA-seq, tRF&tiRNA PCR array, tRNA Repertoire PCR array and tRNA modification analysis.
Promo: 20% OFF! Valid through 7/31/2026
| Service Name | Price |
|---|---|
| tRNA Sequencing Service |
Transfer RNAs (tRNAs) are ubiquitous and the most abundant of all small non-coding RNA molecules. As a fundamental component in translation, tRNAs serve as the physical link be-tween the mRNA coding and protein sequences. A wide variety of biological processes, such as cell proliferation[1], differentiation[1, 2] and apoptosis[3], are always accompanied with variation of tRNAs levels. Alterations of tRNA repertoire affect cell-fate choices during cell development (Fig. 1). Dysregulated tRNA repertoire can promote tumorigenesis and cancer progression [2, 4-11]. Additionally, various diseases show disruptions to the levels and distributions of tRNAs, such as type 2 diabetes mellitus [12], Huntington disease [13], and HIV infection [14]. Studying tRNA repertoire has become an important part of research of biological processes and human diseases.
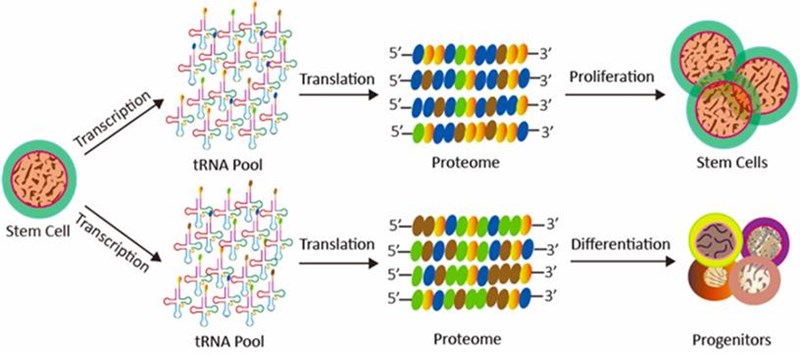
Figure 1. The effects of tRNA repertoire during cell fate determination
tRNAs undergo by far the greatest number of and the most chemically diverse post-transcriptional modifications, which is essential for tRNA stability, folding and decoding. As the amino acid carrier for peptide synthesis, tRNAs must be charged with amino acid. However, these aminoacylated termini and internal modifications impede the adaptor ligation and reverse transcription during tRNA-seq library preparation. Scientists at Arraystar have developed state of the art tRNA-seq methodologies that integrate modification removal and small RNA sequencing optimized for tRNA, ensuring the most reliable and accurate tRNA-seq data for tRNA study.
References
[1] Gingold H, et al. (2014). A dual program for translation regulation in cellular proliferation and differentiation. Cell 158, 1281-92. PMID:25215487.
[2] Pavon-Eternod M, et al. (2013). Overexpression of initiator methionine tRNA leads to global reprogramming of tRNA expression and increased proliferation in human epithelial cells. RNA 19, 461-6. PMID:23431330.
[3] Mei Y, et al. (2010). Apoptotic regulation and tRNA. Protein Cell 1, 795-801. PMID:21113408.
[4] Berns A (2008). A tRNA with oncogenic capacity. Cell 133, 29-30. PMID:18394985.
[5] Waldman YY, et al. (2009). TP53 cancerous mutations exhibit selection for translation efficiency. Cancer Res 69, 8807-13. PMID:19887606.
[6] Kushner JP, et al. (1976). Elevated methionine-tRNA synthetase activity in human colon cancer. Proc Soc Exp Biol Med 153, 273-6. PMID:995958.
[7] Marshall L, et al. (2008). Elevated tRNA(iMet) synthesis can drive cell proliferation and oncogenic transformation. Cell 133, 78-89. PMID:18394991.
[8] Pavon-Eternod M, et al. (2009). tRNA over-expression in breast cancer and functional consequences. Nucleic Acids Res 37, 7268-80. PMID:19783824.
[9] Zhou Y, et al. (2009). High levels of tRNA abundance and alteration of tRNA charging by bor*****ib in multiple myeloma. Biochem Biophys Res Commun 385, 160-4. PMID:19450555.
[10] Begley U, et al. (2013). A human tRNA methyltransferase 9-like protein prevents tumour growth by regulating LIN9 and HIF1-alpha. EMBO Mol Med 5, 366-83. PMID:23381944.
[11] Goodarzi H, et al. (2016). Modulated Expression of Specific tRNAs Drives Gene Expression and Cancer Progression. Cell 165, 1416-27. PMID:27259150.
[12] Krokowski D, et al. (2013). A self-defeating anabolic program leads to beta-cell apoptosis in endoplasmic reticulum stress-induced diabetes via regulation of amino acid flux. J Biol Chem 288, 17202-13. PMID:23645676.
[13] Girstmair H, et al. (2013). Depletion of cognate charged transfer RNA causes translational frameshifting within the expanded CAG stretch in huntingtin. Cell Rep 3, 148-59. PMID:23352662.
[14] van Weringh A, et al. (2011). HIV-1 modulates the tRNA pool to improve translation efficiency. Mol Biol Evol 28, 1827-34. PMID:21216840.
Arraystar tRNA-seq service includes a wealth of bioinformatics analyses, which are valua-ble for investigators to gain insights and better understand the tRNAs in biology and disease.
Differential expression profiling
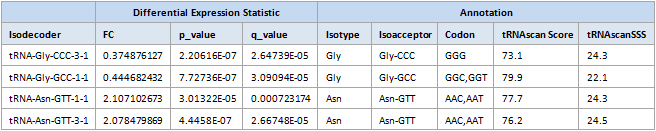

Figure 1. tRNA Expression Profiling is annotated in detail with anticodon, sequence, dot-bracket structure notation, and expression levels.
Plots for differential expression analysis
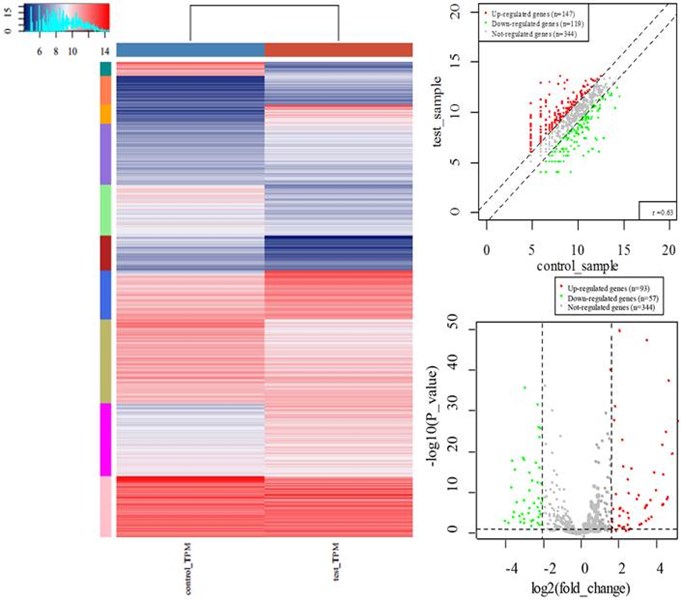
Figure 2. Plots for differential expression analysis. Differentially expressed tRNAs are presented on K-means clustering heatmap, Scatter plot, Volcano plot.
Expression levels of different tRNA types (Additional fee applies)
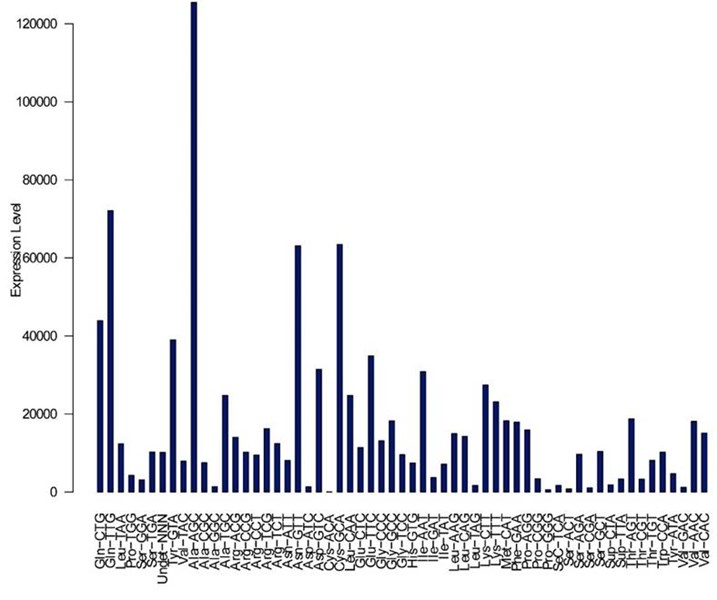
Figure 3. Expression levels of different tRNA types grouped by anticodon.
Threonine fuels glioblastoma through YRDC-mediated codon-biased translational reprogramming. Wu X, et al. Nature Cancer, 2024
Human SAMD9 is a poxvirus-activatable anticodon nuclease inhibiting codon-specific protein synthesis. Zhang F, et al. Science Advances, 2023
Genetic Interaction of tRNA-Dependent Mistranslation with Fused in Sarcoma Protein Aggregates. Lant J T,et al. Genes, 2023
AAV-delivered suppressor tRNA overcomes a nonsense mutation in mice. Wang J, et al. Nature, 2022
Velcrin-induced selective cleavage of tRNALeu(TAA) by SLFN12 causes cancer cell death. Sooncheol Lee, et al. Nature Chemical Biology, 2022
The PPR domain of mitochondrial RNA polymerase is an exoribonuclease required for mtDNA replication in Drosophila melanogaster. Liu Y, et al. Nature Cell Biology, 2022
Neuronal Nsun2 deficiency produces tRNA epitranscriptomic alterations and proteomic shifts impacting synaptic signaling and behavior. Blaze J, et al. Nature Communications, 2021
A rare codon-based translational program of cell proliferation. Guimaraes J C, et al. Genome Biology, 2020
tRNAArg-Derived Fragments Can Serve as Arginine Donors for Protein Arginylation. Avcilar-Kucukgoze I, et al. Cell Chemical Biology, 2020
