Real-Time qPCR Services
MazF-qPCR Service – LncRNA/mRNA/CircRNA
Arraystar MazF qPCR service is based on methyl-sensitive MazF RNase and highly sensitive and accurate qPCR to detect and quantify m6A modification at the exact single nucleotide of the m6ACA site. It can also be used as an independent, non-antibody based method to confirm epitranscriptomic profiling results.
| Service Name | Description | Price |
|---|---|---|
| MazF-qPCR Service - LncRNA/mRNA | Detect and quantify m6ACA sites in LncRNAs/mRNAs | |
| MazF-qPCR Service - Circular RNA | Detect and quantify m6ACA sites in Circular RNAs |
RNase MazF has the enzymatic property to cleave single stranded RNA 5’ immediate to the unmethylated (ACA) sequence, but not methylated (m6ACA) (Fig. 1). RNA treated with MazF and the input RNA without MazF digestion are then quantified by qPCR to determine the extent of m6A modification (Fig. 2).
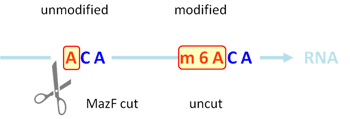
Figure 1. MazF enzyme cuts at unmethylated (ACA) sequence but not methylated (m6ACA).
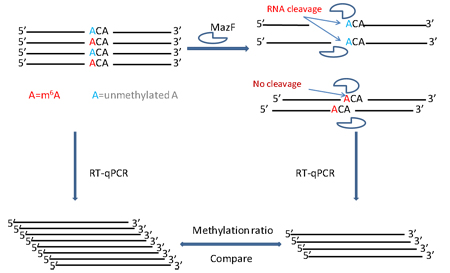
Figure 2. The workflow and procedures of MazF qPCR.
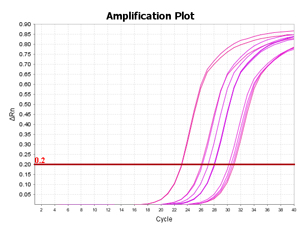
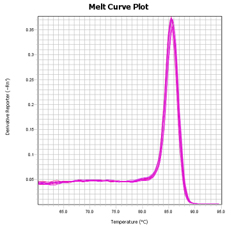
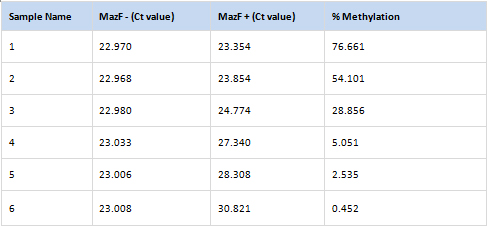
Arraystar MazF qPCR service includes analyses to design primers for quantifiable m6ACA sites, generate amplification plots, post-qPCR melt curve QC, Ct values, and methylation calculation. You send us the total RNA samples, specify the RNA transcripts of interest, and get the m6A modification results.