Related Products
Related Services
Related Reviews
nrStar™ Canonical Conserved miRNA PCR Array (H)
nrStar™ Human Canonical Conserved miRNA PCR Array aims to profile 372 canonical, evolutionarily conserved, functionally significant, mature miRNAs by qPCR. The array efficiently distinguishes the canonical mature miRNAs from their star miRNAs, precursors, and isomiRs using the powerful Two-tailed qPCR technology.
Benefits
• Prioritize the canonical conserved mature miRNAs serving fundamental biology at higher expression levels;
• Focus on the higher expressed, functional mature miRNAs rather than the star miRNAs;
• Highly specifically distinguish a miRNA from its precursor, isomiRs, and family members using the powerful Two-tailed qPCR technology;
• Comprehensive annotation on the miRNA gene promoters, primary miRNAs and host genes;
• Quantify low input samples by powerful preamplification.
The companion rtStar™ First-Strand cDNA Synthesis Kit (3’and 5’adaptor) (Cat # AS-FS-003) is required for the use with nrStar™ Human Canonical Conserved miRNA PCR Array to achieve the best specificity and to distinguish from the precursors and isomiRs. For the low-input samples (100pg ~ 200ng total RNA), rtStar™ PreAMP cDNA Synthesis Kit (Cat#: AS-FS-006) is required for cDNA synthesis and preamplification.
| Product Name | Catalog No. | Description | Size | Price |
|---|---|---|---|---|
| nrStar™ Human Canonical Conserved miRNA PCR Array | AS-NR-005H-1 | 384-well/ plate | $1.00 | |
| nrStar™ Human Canonical Conserved miRNA PCR Array (Roche Light Cycler 480) | AS-NR-005H-1-R | 384-well/ plate | $368.00 |
Conserved Canonical miRNA Collection
Among the 1,917 human miRBase entries, less than a third are confirmed as authentic miRNAs. It is reasonable to prioritize the study on canonical conserved mature miRNAs that are authentic and functionally significant. Therefore, nrStar™ Canonical Conserved miRNA PCR Array is designed to focus on 372 canonical conserved mature miRNAs, offering the researchers a more simple, accurate and targeted solution for miRNA detection.

Figure 1. The Rigorous collection pipeline of the Conserved Canonical miRNAs on the array. All of pre-miRNAs in miRBase are analyzed for canonical pre-miRNAs produced by canonical microRNA biogenesis. These canonical pre-miRNAs are analyzed for phylogenetic conservation to identify the conserved canonical miRNAs. Finally, the higher expressed strand of mature miRNAs over the star strand are selected.
Focus on canonical miRNAs
Only 16% of the 7,095 metazoan miRBase entries are robustly supported as miRNA genes[1]. Canonical miRNAs have broadly distributed expression and yet simultaneously exhibit differential expression among the sample types (Fig 2)[2]. In contrast, most of noncanonical miRNAs are both poorly conserved and lowly expressed in animals, which suggests their lack of significant regulatory functions[3].
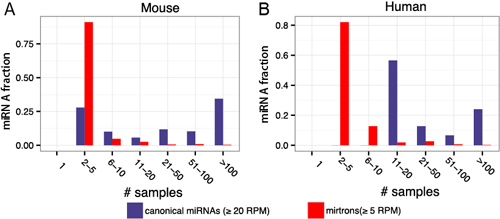
Figure 2. Distribution of canonical miRNAs and mirtrons across small RNA-seq samples.
Evolutionarily conserved
Comparing with newer, less conserved miRNA genes, conserved ones tend to express at higher levels[3, 4]. During the course of evolution, newly emerged miRNA genes were likely to be effectively neutral or deleterious, with large proportions of which were rapidly lost [5]. The conserved miRNAs, on the other hand, have undergone positive selection for their biological roles. In fact, conserved miRNAs have far more conserved mRNA targets than the novel ones, with a high proportion of the mRNA targets encoding transcription factors (Fig 3)[6-8].
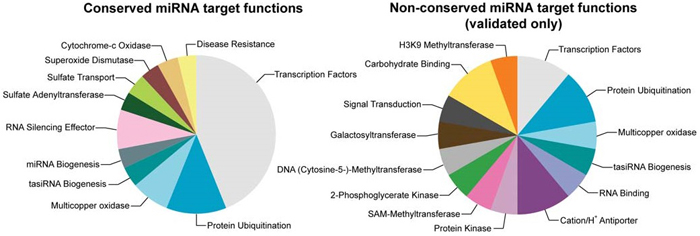
Figure 3. The functional distribution of mRNAs targeted by conserved (left) and nonconserved miRNAs (right), showing prominent enrichment of transcription factor function by canonical miRNAs.
Concentrate on mature miRNAs over star miRNAs
A pre-miRNA is processed into a guide strand and a passenger strand. When the difference between the expression of the two strands is more than 2-fold, the higher expressed strand is designated as mature miRNA and the other as star miRNA in the uniform system for miRNA annotation [1]. If the two strands are similar in amount, both strands are co-mature miRNAs [1]. Star miRNAs tend to be functionally less significant. During evolution, star miRNAs have cumulated a higher nucleotide substitution rate than mature miRNAs, with the seed regions reaching an intriguingly high mutation rate up to 46-fold (Fig 4).
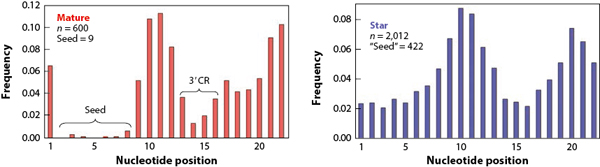
Figure 4. The rate of nucleotide substitution from positions 1 to 22 of mature (colored red on the left) and star miRNAs (colored blue on the right). n, the total number of tallied mutations.
The Powerful Two-tailed miRNA PCR
Due to the small miRNA sizes, the high degree of the homology among family members and isomiRs, and the low abundance in body fluids, miRNA expression profiling is technically challenging. Although qPCR has been used as the “gold standard” for miRNA profiling, traditional qPCR assays rely on the primer specificity for the variations at or near the 3’-end of the miRNA [9, 10], with the 5’-end variations mostly undetected. However, some miRNA family members or isomiRs have only a single nucleotide variation at the 5’-end where the seed region is located. To overcome these challenges, Arraystar has developed a powerful Two-tailed qPCR by elongating the miRNA at both ends to highly specifically interrogate the miRNAs in both 5’- and 3’-regions. The technology has the power to efficiently resolve a miRNA from its precursor, all other family members, and isomiRs, even with low input RNA samples (low to 100 pg total RNA) (Fig 1).
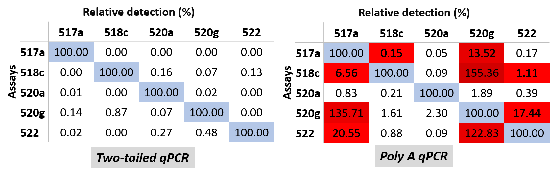
Figure 1. Cross-reactivities as pairwise relative miRNA levels among synthetic miR-515 family members measured by Two-tailed qPCR (left) and PolyA qPCR (right).
Comprehensive annotation
Annotation | Implications |
Tissue-specific | The miRNAs can have the tissue-specific functions in the corresponding tissue. |
Cell-specific | Cell-type specific miRNAs tend to target proteins at crucial bottleneck positions in the corresponding regulatory networks. |
Detection in Serum/plasma | miRNAs in serum/plasma having good potentials in biomarker applications. |
Promoter name | Information about the miRNA biogenesis. Also, many miRNA host genes are associated with the target gene activity either synergistically or antagonistically. |
Promoter Annotation | |
Primary miRNA | |
Host Gene | |
Intronic/intergenic | |
Seed family | miRNAs in the same seed family are evolutionarily closer and have at least partially redundant functions. |
Clustered miRNAs | miRNAs clustered in the genome generally have a similar expression level. They may regulate overlapping sets of target genes through convergent evolution. |
| nrStar™ Human Canonical Conserved miRNA PCR Array (372 miRNAs) |
| Very Broadly conserved miRs (6): miR-31-5p, miR-96-5p, miR-182-5p, miR-184, miR-210-3p, miR-375-3p Broadly conserved miRs (162): let-7a-5p, let-7b-5p, let-7c-5p, let-7d-5p, let-7e-5p, let-7f-5p, let-7g-5p, let-7i-5p, miR-1-3p, miR-7-5p, miR-9-5p, miR-10a-5p, miR-10b-5p, miR-15a-5p, miR-15b-5p, miR-16-5p, miR-17-5p, miR-18a-5p, miR-18b-5p, miR-19a-3p, miR-19b-3p, miR-20a-5p, miR-20b-5p, miR-21-5p, miR-22-3p, miR-23a-3p, miR-23b-3p, miR-24-3p, miR-25-3p, miR-26a-5p, miR-26b-5p, miR-27a-3p, miR-27b-3p, miR-29a-3p, miR-29b-3p, miR-29c-3p, miR-30a-5p, miR-30b-5p, miR-30c-5p, miR-30d-5p, miR-30e-5p, miR-32-5p, miR-33a-5p, miR-33b-5p, miR-34a-5p, miR-34b-5p, miR-34c-5p, miR-92a-3p, miR-92b-3p, miR-93-5p, miR-98-5p, miR-99a-5p, miR-99b-5p, miR-100-5p, miR-101-3p, miR-103a-3p, miR-106a-5p, miR-106b-3p, miR-106b-5p, miR-107, miR-122-5p, miR-125a-5p, miR-125b-5p, miR-126-3p, miR-126-5p, miR-128-3p, miR-129-5p, miR-130a-3p, miR-130b-3p, miR-132-3p, miR-133a-3p, miR-133b, miR-135a-5p, miR-135b-5p, miR-137-3p, miR-138-5p, miR-139-5p, miR-140-3p, miR-141-3p, miR-142-3p, miR-142-5p, miR-143-3p, miR-144-5p, miR-145-5p, miR-146a-5p, miR-146b-5p, miR-147b-3p, miR-148a-3p, miR-148b-3p, miR-150-5p, miR-152-3p, miR-153-3p, miR-155-5p, miR-181a-5p, miR-181b-5p, miR-181c-5p, miR-181d-5p, miR-183-5p, miR-187-3p, miR-190a-5p, miR-191-5p, miR-192-5p, miR-193a-5p, miR-193b-3p, miR-193b-5p, miR-194-5p, miR-195-5p, miR-196a-5p, miR-196b-5p, miR-199a/b-3p, miR-200a-3p, miR-200b-3p, miR-200c-3p, miR-202-3p, miR-202-5p, miR-203a-3p, miR-204-5p, miR-205-5p, miR-206, miR-208a-3p, miR-208b-3p, miR-211-5p, miR-212-5p, miR-214-3p, miR-215-5p, miR-216b-5p, miR-217-5p, miR-218-5p, miR-219a-2-3p, miR-219a-5p, miR-221-3p, miR-222-3p, miR-223-3p, miR-301a-3p, miR-301b-3p, miR-302a-3p, miR-302b-3p, miR-302c-3p, miR-302d-3p, miR-338-3p, miR-363-3p, miR-365a-3p, miR-383-5p, miR-424-5p, miR-425-5p, miR-429, miR-449a, miR-449b-5p, miR-449c-5p, miR-451a, miR-454-3p, miR-455-3p, miR-455-5p, miR-489-3p, miR-490-3p, miR-497-5p, miR-499a-5p, miR-503-5p, miR-551a, miR-551b-3p, miR-802, miR-875-5p Conserved miRs (204): miR-28-3p, miR-105-5p, miR-127-3p, miR-134-5p, miR-136-3p, miR-149-5p, miR-151a-3p, miR-151a-5p, miR-154-3p, miR-154-5p, miR-185-5p, miR-186-5p, miR-188-5p, miR-197-3p, miR-224-5p, miR-296-5p, miR-299-3p, miR-299-5p, miR-323a-3p, miR-323b-3p, miR-324-5p, miR-326, miR-329-3p, miR-330-3p, miR-331-3p, miR-335-5p, miR-337-3p, miR-339-3p, miR-339-5p, miR-340-5p, miR-342-3p, miR-345-5p, miR-346, miR-361-3p, miR-361-5p, miR-362-5p, miR-369-3p, miR-369-5p, miR-370-3p, miR-371a-5p, miR-372-3p, miR-373-3p, miR-374a-3p, miR-374a-5p, miR-374b-5p, miR-376a-3p, miR-376b-3p, miR-376c-3p, miR-377-3p, miR-377-5p, miR-378a-3p, miR-379-5p, miR-380-3p, miR-380-5p, miR-381-3p, miR-382-3p, miR-382-5p, miR-409-3p, miR-410-3p, miR-411-5p, miR-412-5p, miR-421, miR-423-3p, miR-423-5p, miR-431-5p, miR-432-5p, miR-433-3p, miR-448, miR-450a-5p, miR-450b-5p, miR-452-5p, miR-483-5p, miR-485-5p, miR-486-5p, miR-487a-5p, miR-487b-3p, miR-488-3p, miR-488-5p, miR-491-5p, miR-493-5p, miR-494-3p, miR-495-3p, miR-496, miR-498-5p, miR-500a-3p, miR-500b-5p, miR-501-3p, miR-502-3p, miR-504-5p, miR-506-3p, miR-506-5p, miR-507, miR-508-3p, miR-510-5p, miR-511-3p, miR-511-5p, miR-513b-5p, miR-514a-3p, miR-514b-5p, miR-516b-5p, miR-518b, miR-518c-3p, miR-518d-5p, miR-518f-3p, miR-519c-3p, miR-520a-3p, miR-520d-3p, miR-522-3p, miR-523-3p, miR-524-3p, miR-524-5p, miR-525-3p, miR-525-5p, miR-532-5p, miR-539-3p, miR-541-5p, miR-542-3p, miR-543, miR-545-5p, miR-574-3p, miR-576-5p, miR-577, miR-579-5p, miR-582-3p, miR-584-5p, miR-589-5p, miR-590-3p, miR-597-3p, miR-598-3p, miR-605-3p, miR-624-5p, miR-628-5p, miR-651-5p, miR-652-3p, miR-654-3p, miR-656-3p, miR-660-5p, miR-671-3p, miR-671-5p, miR-675-3p, miR-676-3p, miR-708-5p, miR-744-5p, miR-758-3p, miR-760, miR-769-5p, miR-770-5p, miR-873-3p, miR-873-5p, miR-874-3p, miR-876-3p, miR-885-5p, miR-887-3p, miR-887-5p, miR-888-5p, miR-889-3p, miR-890, miR-892a, miR-892b, miR-892c-3p, miR-934, miR-935, miR-944, miR-1180-3p, miR-1185-1-3p, miR-1185-2-3p, miR-1185-5p, miR-1197, miR-1247-5p, miR-1249-3p, miR-1251-5p, miR-1264, miR-1271-5p, miR-1277-5p, miR-1283, miR-1298-5p, miR-1301-3p, miR-1307-3p, miR-1307-5p, miR-1323, miR-1343-3p, miR-1911-5p, miR-1912-3p, miR-2114-3p, miR-2114-5p, miR-2355-3p, miR-2681-3p, miR-2681-5p, miR-3059-5p, miR-3085-3p, miR-3140-3p, miR-3145-3p, miR-3146, miR-3173-5p, miR-3200-3p, miR-3613-5p, miR-3617-5p, miR-3934-5p, miR-4524a-3p, miR-4677-3p, miR-4766-3p, miR-4766-5p, miR-6529-5p, miR-9851-3p |
Compatible qPCR Instruments Equipped with a 384-well Format Block:
ABI ViiA™ 7
ABI 7500 & ABI 7500 FAST
ABI 7900HT
ABI QuantStudio™ 5 Real-Time PCR system
ABI QuantStudio™ 6 Flex Real-Time PCR system
ABI QuantStudio™ 7 Flex Real-Time PCR system
ABI QuantStudio™ 12K Flex Real-Time PCR System
Bio-Rad CFX384
Bio-Rad iCycler & iQ Real-Time PCR Systems
Eppendorf Realplex
Roche LightCycler 480
Stratagene Mx3000

