Real-Time qPCR Services
Related Products
Related Services
Related Reviews
MeRIP-qPCR Service – LncRNA/mRNA/CircRNA
Arraystar MeRIP-qPCR service combines methylated RNA immunoprecipitation (MeRIP) with highly sensitive and accurate qPCR to detect and quantify m6A modification in a whole transcript/gene or fragment.
| Service Name | Description | Price |
|---|---|---|
| MeRIP-qPCR Service - LncRNA/mRNA | Quantify m6A modification level in mRNAs/LncRNAs | |
| MeRIP-qPCR Service - Circular RNA | Quantify m6A modification level in Circular RNAs |
RNA samples are either unfragmented (intact) for any modification sites in a whole transcript/gene, or fragmented for specific m6A sites of interest. The RNAs are immunoprecipitated with a highly specific antibody against the modification (e.g. m6A). IP enriched RNA and the input RNA are then quantified by qPCR to determine the extent of m6A modification in the entire RNA transcript/gene or at the interrogated methylation site (Fig. 1).
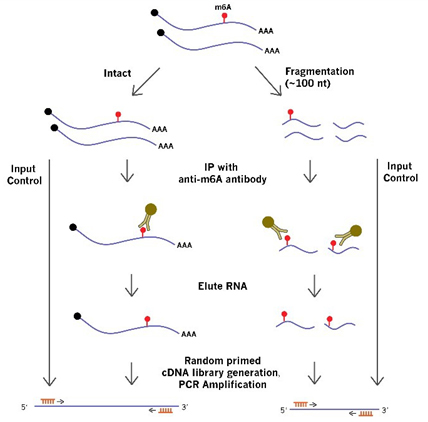
Figure 1. The workflow and procedures of MeRIP-qPCR.
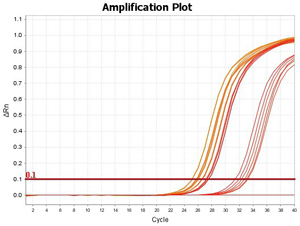
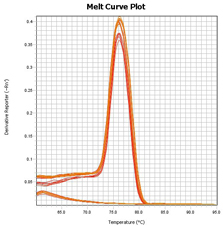
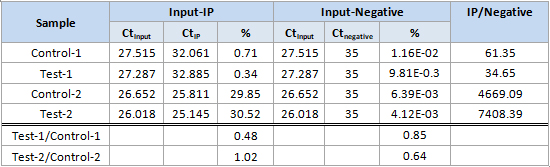
Arraystar MeRIP-qPCR service includes MeRIP experiment, primer design, amplification plots, post-qPCR melt curve QC, Ct values, and methylation calculation. You send us the total RNA samples, specify the genes and methylation sites of interest, and get the m6A modification results.
N6- methyladenosine modification of a parvovirus- encoded small noncoding RNA facilitates viral DNA replication through recruiting Y- family DNA polymerases. Ning K, et al. Proceedings of the National Academy of Sciences of the United States of America, 2024
