NGS Solutions
Related Products
Related Services
Related Reviews
eccDNA Sequencing Service
In eccDNA sequencing, the overwhelming amounts of linear chromosomal DNA and circular mitochondrial DNA (mtDNA) in the DNA sample must be removed as they can significantly reduce the eccDNA reads. Arraystar eccDNA-seq linearizes the circular mtDNAs by restriction endonuclease cleavage and then digests the linearized mtDNA and linear chromosomal DNA by a linear DNA-specific exonuclease, leaving eccDNAs intact and enriched in the sample. The eccDNAs are amplified by Rolling Circle Amplification (RCA), greatly improving the eccDNA signals and data quality.
Benefits
• Broad coverage: Simultaneous profiling of eccDNA(<10 kb) and ecDNA(>=10 kb).
• Maximal eccDNA enrichment: Removal of chromosomal and mitochondrial DNA and Rolling Circle Amplification of eccDNA to dramatically increase eccDNA reads.
• High reliability: Well established and optimized procedures to produce best possible results.
• Easy-to-use analyses: Provided with rich annotation, genome browser track visualization, publication-ready graphics for biologists and clinicians.
Watch Video> The Extraordinary eccDNAs in Cancer and Diseases
Promo: 20% OFF! Valid through 6/30/2026
| Service Name | Price |
|---|---|
| eccDNA Sequencing Service |
Extrachromosomal circular DNA (eccDNA) is a class of double-stranded circular DNA molecules present outside of regular chromosomal DNA. eccDNA is widely found in human normal and cancer tissues, body fluids, and other eukaryotic cells. According to the lengths, eccDNA can be classified into two main categories : eccDNA (<10 kb) and ecDNA (>=10 kb)[1]. eccDNAs may arise from any regions of the genome in normal tissues or as a byproduct of programmed cell death. These entities are typically devoid of genes and are not subjected to either amplification or growth selection within the cells. The term extrachromosomal DNA (ecDNA) is more commonly used to refer specifically to the very large eccDNAs, i.e. large clonal circular DNA molecules (10 kb to several Mb) that are clonally inherited, self-replicating, amplifying and selected only in cancer cells. These ecDNA molecules frequently harbor oncogenes, drug resistance genes, or mobile super-enhancers (Fig. 3), which provide an evolutionary selective advantage for cancers (Fig. 1). Functionally, eccDNAs and ecDNAs regulate diverse molecular and physiological processes (Fig. 2) [2], are closely associated with a variety of diseases (Table 1), and can be excellent diagnostic and prognostic markers (Table 2). In recent years, eccDNAs have become one of the major breakthroughs and research hotspots in cancer research.
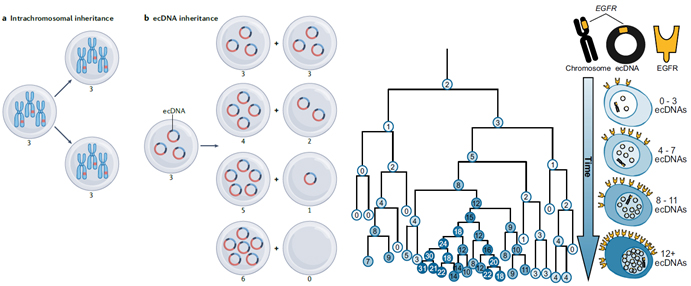
Figure 1. The uneven segregation of eccDNAs (left, b) promotes tumor genetic heterogeneity (right), which provides a strong evolutionary advantage for cancer cells [3, 4].
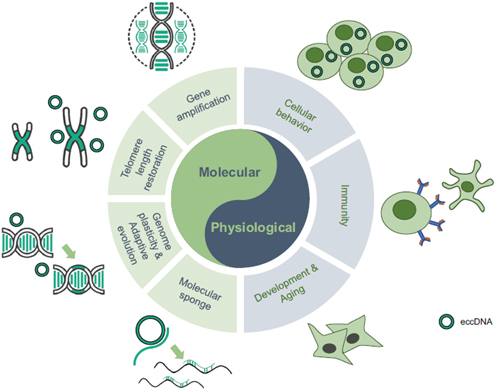
Figure 2. eccDNAs regulate diverse molecular and physiological processes. Mechanistically, eccDNAs play important roles in gene amplification, telomere length restoration, genomic plasticity and molecular sponges, involved in a variety of physiological events, e.g. cellular behavior, immunity and aging [2].
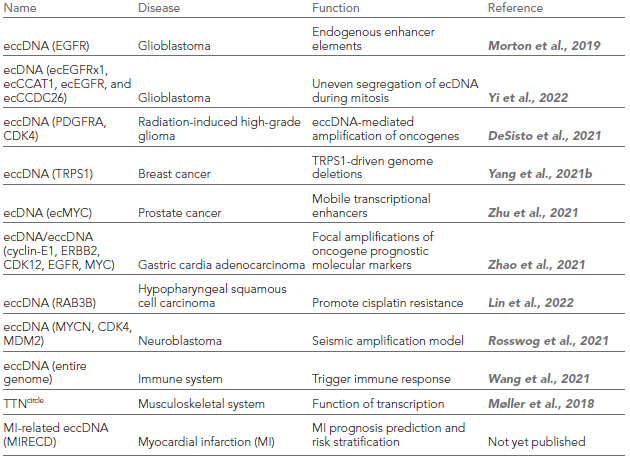
Table 1. eccDNAs associated diseases [5].
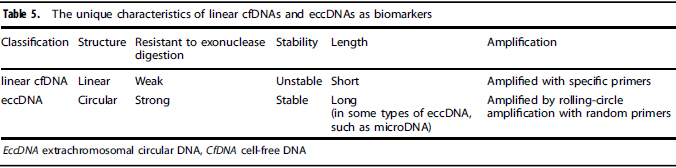
Table 2. The unique advantages of eccDNA over linear cfDNA as biomarkers [2]
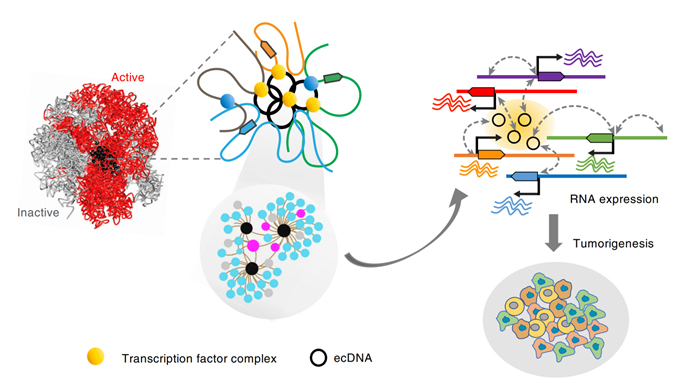
Figure 3. ecDNA acts as a mobile super-enhancer to regulate gene transcription [6].
References
[1] Hung, K. L., et al. (2022) “Gene regulation on extrachromosomal DNA” Nat Struct Mol Biol 29(8):736-744 [PMID: 35948767]
[2] Yang, L., et al. (2022) “Extrachromosomal circular DNA: biogenesis, structure, functions and diseases” Signal Transduct Target Ther 7(1):342 [PMID: 36184613]
[3] Noorani, I., et al. (2022) “Leveraging extrachromosomal DNA to fine-tune trials of targeted therapy for glioblastoma: opportunities and challenges” Nat Rev Clin Oncol 19(11):733-743 [PMID: 36131011]
[4] Nathanson, D. A., et al. (2014) “Targeted therapy resistance mediated by dynamic regulation of extrachromosomal mutant EGFR DNA” Science 343(6166):72-6 [PMID: 24310612]
[5] Zhao, Y., et al. (2022) “Extrachromosomal circular DNA: Current status and future prospects” Elife 11([PMID: 36256570]
[6] Zhu, Y., et al. (2021) “Oncogenic extrachromosomal DNA functions as mobile enhancers to globally amplify chromosomal transcription” Cancer Cell 39(5):694-707 e7 [PMID: 33836152]

Fig 1. The workflow of eccDNA Sequencing
1. Genomic DNA extraction
2. Linearization of mitochondrial DNA
3. Removal of linear DNA
4. Rolling cycle amplification
5. Ultrasound fragmentation and DNA sequencing library construction
6. High-throughput sequencing
7. Data analysis and report
8. Delivered results
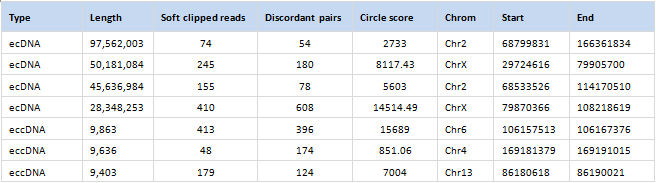
Annotated eccDNA data table
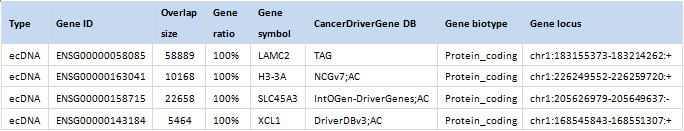
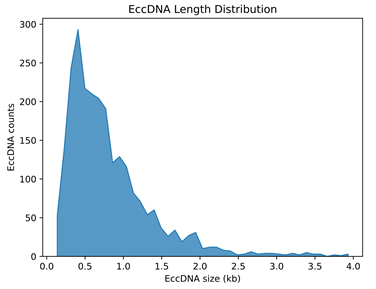
Histogram of eccDNA size distribution
Distribution of eccDNAs across the genome

