The Challenges and Solutions of Small RNA Profiling
The Biases of Using TPM in Small RNA Sequencing Data Analysis
Simultaneously Profile Multiple Small RNA Classes Accurately
Direct End-labeling to Avoid the Biases From Sequencing Library Prep
Mature small RNAs like miRNAs and tsRNAs have non-random, precisely cleaved termini and defined RNA sizes during biogenesis. Arraystar Small RNA Microarray probes are designed to maximize the sensitivity and specificity for both sequence and size discrimination for even closely related target RNAs.
For mature small RNA probes (Fig.1), the 5’-CAP hairpin structure and the fixed G position for C-Cy3 label provide much enhanced discriminating power for the mature small RNA size and terminal end. The short Specific Probe region matches to the optimal probe/target Tm for the hybridization condition. Along with the proclivity of small RNA precursors to form much more stable intra-molecular secondary structure and the relatively lower precursor RNA levels, the mature small RNA probes have excellent distinct specificity to differentiate mature vs precursor small RNA signals.
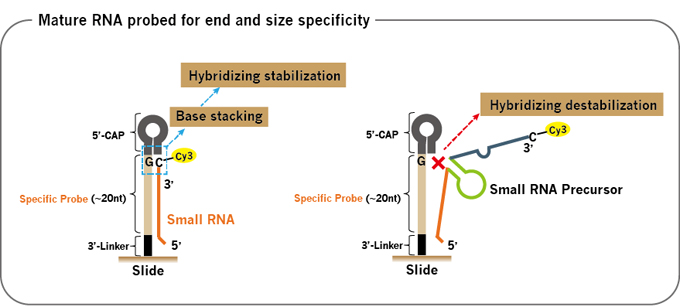
Figure 1. Mature small RNA probe. From 5’->3’, (A) The 5′-Cap segment forms a hairpin stem structure to destabilize longer size sequences such as precursors, isoforms from annealing. (B) The fixed G at the first base position in the probe hybridization region base pairs with the C-Cy3 label at the 3’-end of the target RNA, forming more stable GC-clamp. (C) The short ~20-nt Specific Probe region is normalized for optimal Tm to distinguish similar RNAs with only 1~2 base differences. (D) 3’-Linker provides the space to prevent steric hindrance from the slide surface.
For precursor/longer small RNA probes (Fig.2), the probes are designed to specifically hybridize the unique sequences of the pre-miRNAs, tRNAs, and snoRNAs.
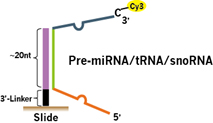
Figure 2. Small RNA precursor probe. The ~20-nt short probe binding region is Tm optimized with the unique sequences of the precursor pre-miRNA, tRNA and snoRNA.
For the collections of miRNAs, pre-miRNAs, tsRNAs, tRNAs, and snoRNAs on the microarray, the probe sets accurately detect and quantify these small RNAs even with only one or two base differences.
Related Service
