The Challenges and Solutions of Small RNA Profiling
The Biases of Using TPM in Small RNA Sequencing Data Analysis
Simultaneously Profile Multiple Small RNA Classes Accurately
Direct End-labeling to Avoid the Biases From Sequencing Library Prep
To fluorescently label small RNAs for microarray profiling, the total RNA is treated with T4 polynucleotide kinase (T4 PNK) to remove both monophosphate (P) and cyclic phosphate (cP) groups from the 3′ end to expose the 3-OH end. DMSO is added as a denaturant to minimize the effects of structure and sequence context differences among small RNAs. The small RNAs are then ligated with pCp-Cy3 onto the 3’-ends by T4 RNA ligase. One RNA molecule is labeled with one Cy3 label (Fig. 1).
Advantages
- Eliminates biases from cDNA synthesis by reverse transcription due to RNA modification interference and RNA folding hindrance as in small RNA-seq.
- Avoids distortions from PCR amplification cycles as in required small RNA-seq library amplification.
- Uses DMSO to reduce the RNA structure and sequence context differences among small RNAs.
- Simple and straightforward procedures, requiring much lower total RNA amounts (100ng) compared to small RNA-seq. No complex procedures, e.g. small RNA size fractionation or multiple purification steps that cause sample losses.
- More tolerant of internal RNA chemical damages, modifications, or degradations than polymerase-based methods. Particularly advantageous for archived or chemically preserved samples such as FFPE RNAs.
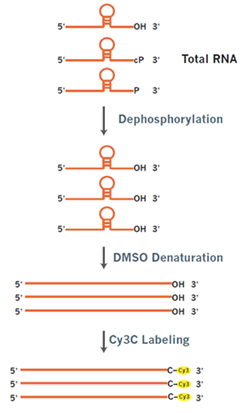
Figure 1. Direct end-labeling of Small RNA. RNAs with 3’-P or 3’-cP ends are dephosphorylated with T4 polynucleotide kinase (T4 PNK) to form 3-OH ends. DMSO is added to reduce the effects of RNA structures on the labeling. pCp-Cy3 is ligated to the 3’-OH of the RNA by T4 RNA ligase. One RNA is terminally labeled with exactly one cyanine dye label.
Related Service
