The tRNA pool, composed of tRNA isoacceptor families for each amino acid, is an important source of insider information in the study of codon usage, protein translation efficiency and accuracy, biological processes and human diseases. A codon may be decoded by a perfectly matched cognate anticodon, or, by a near-cognate tRNA with a single mismatch at the wobble position. A cognate tRNA supply exceeding protein translation demands is the optimal codon for translation. The cognate:near-cognate tRNA ratios in the tRNA pool are dynamically regulated to affect translation efficiency, fidelity, and transcript stability (Sup. Fig. 1)[1]. Under various conditions, tRNA pools can alter in ways to maintain homeostasis or to favor a particular gene expression program [2]. The perturbed levels of isoacceptors can lead to diseases such as cancer (Sup. Fig. 2)[3] and neurodegeneration (Fig.1)[4].
 |
|
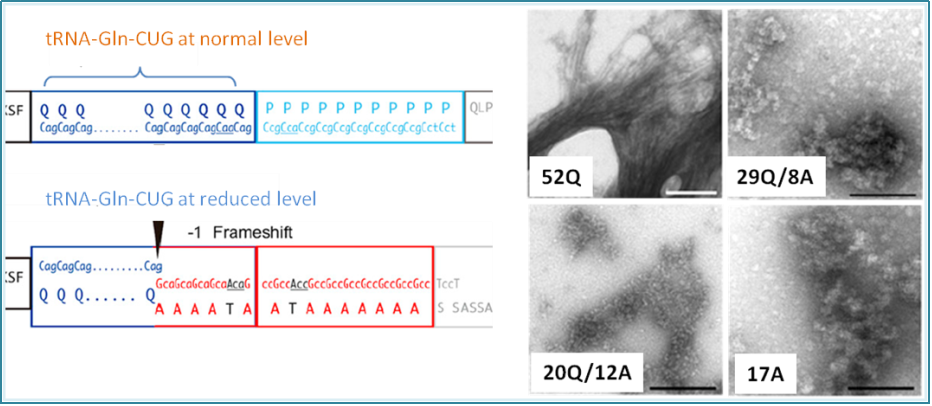
Figure 1. tRNA-Gln-CUG at normal supply level correctly translates poly-Q repeat region in Huntintin. Reduced tRNA-Gln-CUG level causes -1 frame shift translation to make erroneous poly-Alanines. The extent of Q/A ratio in Huntingtin determines the aggregation state (from normal 52Q to pathogenic 17A) and thus the severity of Huntington’s disease [4]. |
nrStar™ tRNA PCR Arrays profile 185 Human/Mouse tRNAs, covering all the nuclear anticodon isoacceptors catalogued in GtRNAdb, providing insights on both the isoacceptor and the specific isodecoder expression.
To learn more about how tRNAs become active regulators, watch our webinar now!
References
[1] Hanson G. and J. Coller (2018) Nat. Rev. Mol. Cell Biol. [PMID: 29018283]
[2] Gavin H. et al. (2014) Molecular Cell Biology [PMID: 29018283]
[3] Goodarzi H. et al. (2016) Cell [PMID: 27259150]
[4] Girstmair H. et al. (2013) Cell Rep [PMID: 23352662]
|

