How to Choose the Appropriate Tool for Small RNA Modification Research?
Molecular Mechanisms of Small RNA Modifications
Small RNA Modifications: Integral to Functions and Diseases
The Challenges and Solutions of Studying Modified Small RNAs
Small RNAs are modified by chemical groups that regulate the RNA activities or endow them with new functions. By various molecular mechanisms, for example, the modifications can alter the miRNA targeting specificities or change the tsRNA binding affinities with RNA binding proteins to exert biological activities. Profiling small RNA modifications is a new frontier of epitranscriptomic research of great scientific and clinical importance.
Modified small RNAs redirect target repression
Modifications in the seed regions (positions 2–8) of miRNAs can change the miRNA repression targets. For example, O8G modified miR-184 redirects it targeting specificity toward otherwise mismatched Bcl-xL and Bcl-w and suppresses their translation (Fig.1) [1]. In another example, upon treatment with an adrenergic agonist, increased O8G modified miR-1 redirects to a new set of targets that function in cardiac hypertrophy pathways [2]. In fact, under pathophysiological conditions, O8G modification of miRNAs is a general epitranscriptional mechanism as a part of redox signaling by which ROS imparts fine tuning of gene expression [2, 3].
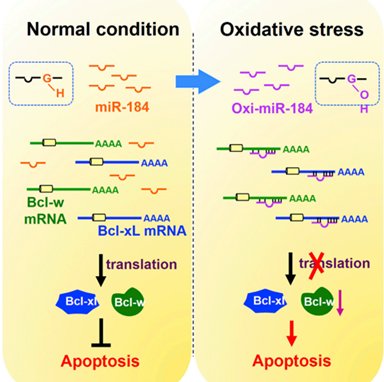
Figure 1. O8G-modified miR-184 redirects targeting activity at mismatched Bcl-xL and Bcl-w mRNAs and suppresses their anti-apoptosis protein translation, which leads to increased susceptibility of the heart to myocardial ischemia/reperfusion (I/R) injury.
Modified small RNAs decoy RNA binding proteins (RBP)
Modified small RNAs can decoy RNA binding proteins (RBP), thereby inhibiting RBP binding to target mRNAs and modulating gene expression by modification-dependent regulation. For example, the 5’ terminal oligoguanine (TOG) in tsRNAs can either have an unmodified uridine (U8) or pseudouridine (Ψ8) modified by PUS7 at the 8th position. The U8/Ψ8 status determines the different binding affinities with polyadenylate-binding protein 1 (PABPC1) as a translational initiation factor. Ψ8-TOG-5’tsRNAs bind more tightly to PABPC1, displace it from mRNAs, and inhibit the translation, whereas U8-TOG-5’ tsRNAs do not (Fig. 2)[4].
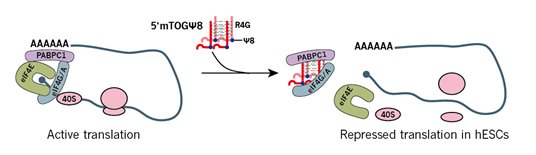
Figure 2. 5’TOG-tsRNAs with Ψ modification at position 8 decoy and displace PABPC1 from the translation initiation complex, thus repressing the translation of capped mRNAs [4].
Related Service
Small RNA Modification Array Service
References
[1] Wang, J. X., et al. (2015) “Oxidative Modification of miR-184 Enables It to Target Bcl-xL and Bcl-w” Mol Cell 59(1):50-61 [PMID: 26028536]
[2] Seok, H., et al. (2020) “Position-specific oxidation of miR-1 encodes cardiac hypertrophy” Nature 584(7820):279-285 [PMID: 32760005]
[3] Seok, H., et al. (2016) “MicroRNA Target Recognition: Insights from Transcriptome-Wide Non-Canonical Interactions” Mol Cells 39(5):375-81 [PMID: 27117456]
[4] Guzzi, N., et al. (2018) “Pseudouridylation of tRNA-Derived Fragments Steers Translational Control in Stem Cells” Cell 173(5):1204-1216 e26 [PMID: 29628141]
