Introduction
R-loops are three-stranded RNA:DNA loops when the nascently transcribed RNA anneals with the template DNA strand and displaces the non-template DNA as unpaired single strand. Besides nascent mRNAs, lncRNAs or circRNAs may also create and form R-loops. Mounting evidences now support cells can harness R-loops to regulate gene expression in ways including epigenetic regulation, transcriptional initiation and elongation. R- loop misregulation is associated with DNA damage, hyper-recombination, and genomic instability.
mRNA organized R-loops
• DNA methylation regulation by mRNA organized R-loop
R-loop accumulation is a characteristic feature of unmethylated CGI-containing promoters that also show GC skew[1]. In genome-wide survey, R-loops are enriched at loci with decreased DNA methylation, increased DNase hypersensitivity, and higher chromatin accessibility[2]. R-loops can inhibit DNA methylation by repelling the binding of 5mC DNA methyltransferases (DNMTs)(Fig.1)[3,4].

Figure 1. Normally in healthy cells, The R-loop at gene promoter repels DNMT binding, prevents DNA methylation, and facilitates transcription. However in amyotrophic lateral sclerosis (ALS4), senataxin mutation reduces R-loop formation, leading to increased the negative TGFb regulator BAMBI gene promoter methylation, transcription repression, TGFb signal transduction upregulation, and ALS progression [4].
• mRNA organized R-loop as promoter
mRNA transcription by RNA polymerase II (Pol II) is initiated more efficiently on accessible, nucleosome depleted gene promoters. Similarly, the ssDNA component of R-loops has the potential to directly promote Pol II transcription, without the need for typical promoter double dsDNA unwinding by general transcription factors (GTFs). The R-loops, formed by nascent mRNA invading the DNA duplex behind the progressing PolII, can functionally act as a promoter for divergent antisense LncRNA (AS transcript)(Fig.2). There, the ssDNA in the R-loop becomes the template for the antisense RNA.

Figure 2. R-loop emerged in the gene promoter region during active mRNA transcription can functionally act as a promoter for the divergent antisense lncRNA transcription. S transcript: sense RNA (mRNA); AS transcript: antisense RNA; GTF: General transcription factor; pol II: polymerase II [5].
lncRNA organized R-loops
• DNA methylation regulation by lncRNA organized R-loops
A mechanism whereby R-loops favor DNA hypomethylation is by attracting DNA demethylases (TET). For example, in mESC, antisense lncRNA TARID forms an R-loop at the TCF21 gene promoter, which binds GADD45A protein to recruit TET1, leading to transcriptional activation of TCF21 (Fig.3)[6].
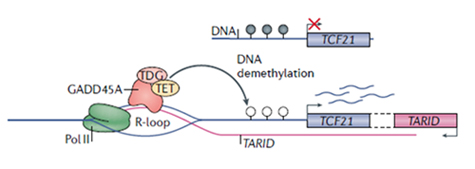
Figure 3. R-loop by antisense lncRNA TARID binds GADD45A, recruits TET1 to the CpG island promoter of TCF21 gene, and demethylates the DNA, activating the TCF21 transcription [6].
• Chromatin remodeling by lncRNA organized R-Loops
In certain cell lines and tissues, lncRNA ANRASSF1 is transcribed from the strand opposite to the RASSF1 mRNA gene. ANRASSF1 endogenous expression is higher in breast and prostate tumor cells compared with non-tumor. The R-loop formed in cis by ANRASSF1 directs PRCs (e.g. SUZ12) to the target RASSF1A gene promoter, selectively repressing the expression of RASSF1-A isoform(Fig. 4)[7].

Figure 4. Antisense lncRNA ANRASSF1 forms R-loop in the RASSF1A promoter region, recruiting PRC2 chromatin modifier to silence RASSF1A expression via repressive H3K27me3 histone mark [7].
• R-loop as lncRNA anchor
In this case, an R-loop only serves as the anchor for tethering the lncRNA to the specific DNA site. It is the tethered single-stranded lncRNA that recruits chromatin remodeling complexes.
Example 1
The bidirectional lncRNA of the CCND1 gene is transcribed as predominantly single-stranded, DNA-bound multiple isoforms. A portion of the lncRNA length is tethered to the R-loop in the 5’ regulatory region of CCND1 gene. The other part of the single-stranded RNA recruits TLS repressosome to the CCND1 promoter to repress the gene (Fig.5)[8].

Figure 5. Nascently transcribed CCND1 lncRNAs are tethered to the R-loops, recruiting TLS repressosome to the CCND1 promoter to negatively regulate CCND1 mRNA transcription.
Example 2
GATA3 mRNA and GATA3-AS1 lncRNA belong to the general class of bidirectional lncRNA/mRNA pairs. Transcriptional start sites (TSS) for GATA3 and GATA3-AS1 are separated by ∼1200 bp. GATA3-AS1 is necessary for efficient transcription of GATA3. A portion of GATA3-AS1 forms the R-Loop and the other part of the same RNA recruits MLLH3K4 methyltransferase to the promoter region of the GATA3 gene, modifies the histone with activating H3K4me3 mark, and promotes GATA3 mRNA transcription [9].

Figure 6. Antisense LncRNA (GATA3-AS1) forms an R-Loop in the shared promoter region of the GATA3 and GATA3-AS1 gene. The single-stranded portion of the GATA3-AS1 tethered to the R-loop recruits MLL methyltransferase to mark the promoter region with activating H3K4me3 and facilitates GATA3 transcription [9].
• lncRNA organized R-loop as regulator of transcription factor/repressor binding
For example, at the VIM locus, the antisense lncRNA VIM-AS1 forms an R-loop in the promoter region of the VIM coding gene, recruiting NF-κB to promote VIM mRNA expression (Fig.7)[10]. Therefore, VIM mRNA levels are positively correlated with the expression of VIM-AS1, with both being silenced in primary colon cancers concomitant with hypermethylation of the promoter.

Figure 7. Antisense lncRNA VIM-AS1 forms an R-loop in the promoter region of VIM coding gene, recruiting NF-κB to promote VIM mRNA expression [10].
• lncRNA organized R-loop stabilization in transcription regulation
Stabilization of R-loops can be a general mechanism to inhibit transcriptional elongation in many organisms. In A. thaliana, lncRNA COOLAIR is a set of antisense transcripts originating from the 3′ end of FLOWERING LOCUS C (FLC), which are induced by cold temperature. The R-loop covering the COOLAIR promoter can be stabilized by homeodomain protein AtNDX via its binding to the ssDNA strand. The stable R-loop can cause Pol II stalling and abortion of COOLAIR transcription, which in turn regulates the FLC expression [11].

Figure 8. The R-loop stabilized by AtNDX binding causes Pol II stalling and abortion of COOLAIR transcription [11].
As another example, human Snord116 locus responsible for Angelman syndrome has the propensity for forming R loops because of the repeating units of GC skew. Under physiologic conditions, the polII transcription machinery is able to progress all the way through to make Ube3a-ATS to silence Ube3a mRNA gene expression in cis (Fig.9). However, under conditions such as topoisomerase inhibitor topotecan treatment, the R-loop formation is increased and stabilized, which causes excessive transcriptional stalling, shutting down of Ube3a-ATS, relief of silencing on Ube3a, and up-regulation of Ube3a mRNA expression [12].


Figure 9. R-loop mechanism of topotecan drug action on ubiquitin ligase Ube3a gene expression and Angelman syndrome. (Top) Schematic of snord116 locus for mRNA Ube3a and antisense RNAs Snrpn, snord116, and Ube3a-ATS. Ube3a-ATS can silence the Ube3a expression by extending into the Ube3a template. The snord116 locus is under the control of imprinting control region (ICR). (Bottom) Topotecan drug inhibition of topoisomerase increases and stabilizes R-loop formation within the snord116 locus, which stalls polII transcriptional progression through Ube3a-ATS. The reduction of antisense Ube3a-ATS relieves the silencing on Ube3a mRNA expression and Angelman syndrome [12].
• Trans-induced lncRNA organized R-loops
Although most R-loops are formed via co-transcription from the same locus in cis, some G-rich trans-acting RNAs can form trans-induced R-loops through continuous threading into G-quadruplexes (G4s) structures by base pairing with the C-rich strand. Whereas cis-induced R loops are restricted to the same sites of RNA transcription, trans-induced R loops can occur across the genome, creating multiple ‘‘hot spots’’ for genomic instability. Therefore, trans-induced R loops could be a bigger threat to genome integrity.
As a trans-induced R-loop example, lncRNA APOLO (auxin-regulated promoter loop) is responsible for the activation of auxin responsive genes in Arabidopsis thaliana. The target genes of APOLO are normally silenced and maintained by the Polycomb factor like heterochromatin protein 1 (LHP1). In response to auxin, the transcribed APOLO forms R-loops by sequence complementarity with multiple distant tans target sites, where it is anchored to decoy away LHP1, allowing the target gene expression [13]. The APOLO sequence has two TTCTTC cores that could perfectly recognize the consensus motif sequence GAAGAA(G/C) in the target genes by sequence complementarity.

Figure 10. Arabidopsis thaliana lncRNA APOLO can form R-loops in trans by sequence complementarity with multiple distant targets as part of a widespread regulation of auxin responsive genes. Modulating APOLO RNA levels by auxin affects R-loop formation and the transcriptional activities of its target loci, including auxin-responsive genes (e.g. WAG2 and AZG2) during lateral root formation [13].
CircRNA organized R-loops
SMARCA5 is an important protein factor in DNA repair. It participates in chromatin remodeling at the DNA damage sites and promotes the recruitment of DNA repair factors. SMARCA5 gene can produce circSMARCA5 circular RNA when the exon16 and exon15 are backspliced. In breast cancer cells, circSMARCA5 expression is significantly down regulated. Overexpression of circSMARCA5 induces drug sensitivity of the breast cancer cells [14]. Mechanistically, circSMARC5 can form R-loop with its host gene DNA. The R-loop pauses the exon15 transcription which causes SMARCA5 protein truncation and dysfunction (Fig. 11).

Figure 11. circSMARCA5 forms R-loop in the SMARCA5 host gene body region, leading to transcription pause, truncated mRNA transcript, defective SMARCA5 protein degradation, and impairment of DNA repair process [14].
Related Service
DRIPc-Sequencing Service
References
1. Ginno PA. et al (2013) GC skew at the 5' and 3' ends of human genes links R-loop formation to epigenetic regulation and transcription termination. Genome Res 23(10):1590-600 [PMID:23868195].
2. Nadel J. et al (2015) RNA:DNA hybrids in the human genome have distinctive nucleotide characteristics, chromatin composition, and transcriptional relationships. Epigenetics Chromatin 8:46 [PMID:26579211].
3. Ginno PA. et al (2012) R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol Cell 45(6):814-25 [PMID:22387027]
4. Grunseich C. et al (2018) Senataxin Mutation Reveals How R-Loops Promote Transcription by Blocking DNA Methylation at Gene Promoters. Mol Cell 69(3):426-437.e7 [PMID:29395064]
5. Tan-Wong SM. et al (2019) R-Loops Promote Antisense Transcription across the Mammalian Genome. Mol Cell 76(4):600-616.e6 [PMID:31679819]
6. Arab K. et al (2019) GADD45A binds R-loops and recruits TET1 to CpG island promoters. Nat Genet 51(2):217-223 [PMID:30617255]
7. Beckedorff FC. et al (2013) The intronic long noncoding RNA ANRASSF1 recruits PRC2 to the RASSF1A promoter, reducing the expression of RASSF1A and increasing cell proliferation. PLoS Genet 9(8):e1003705 [PMID:23990798]
8. Wang X. et al (2008) Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription. Nature 454(7200):126-30 [PMID:18509338]
9. Gibbons HR. et al (2018) Divergent lncRNA GATA3-AS1 Regulates GATA3 Transcription in T-Helper 2 Cells. Front Immunol 9:2512 [PMID:30420860]
10. Boque-Sastre R. et al (2015) Head-to-head antisense transcription and R-loop formation promotes transcriptional activation. Proc Natl Acad Sci U S A 112(18):5785-90 [PMID:25902512]
11. Sun Q. et al (2013) R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 340(6132):619-21 [PMID:23641115]
12. Powell WT. et al (2013) R-loop formation at Snord116 mediates topotecan inhibition of Ube3a-antisense and allele-specific chromatin decondensation. Proc Natl Acad Sci U S A 110(34):13938-43 [PMID:23918391]
13. Ariel F. et al (2020) R-Loop Mediated trans Action of the APOLO Long Noncoding RNA. Mol Cell 77(5):1055-1065.e4 [PMID:31952990]
14. Xu X et al (2020) CircRNA inhibits DNA damage repair by interacting with host gene. Mol Cancer 19(1):128. [PMID:32838810]