Why Is m6A Modification in Circular RNA Important?
CircRNAs are an unusual RNA class exerting their biological functions by regulating gene transcription and splicing [1, 2], encoding small peptides [3, 4], and acting as sponges for miRNAs through their miRNA binding sites [5, 6]. Like in mRNAs, m6A is the most abundant internal epitranscriptomic modification in circRNAs [7]. m6A modification in circRNAs is installed by writers, e.g. METTL3 and METTL14 and is reversibly demethylated by erasers such as FTO and ALKBH5.
m6A modification is functionally significant in circRNA biological activities. Beyond the well-known m6A function in regulating circRNA stability [7], m6A can modulate circRNA-miRNA interactions and consequently the circRNA function as miRNA sponges, affect the interaction between circRNAs and RNA-binding proteins (RBP) [8], and mark the endogenous circRNAs as “self” from host immune response to foreign RNAs [9].
Notably, both modified and unmodified forms of circRNAs are usually found in the same cell at variable ratios. As the modified and unmodified fractions (modification stoichiometry) of the same circRNA can assume different fates, determining the circRNA modification percentage is highly relevant to their functional consequences [10, 11].
m6A regulates circRNA stability
m6A modification in circRNAs can be read by YTH-DF2, which binds HRSP12 adaptor protein and recruits endoribonucleases RNase P/MRP. The endonucleases cleave open the RNA circle, causing rapid degradation of the m6A modified circRNAs (Fig.1) [12].
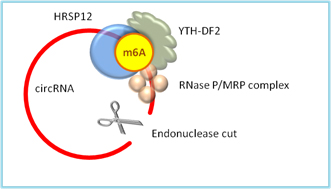
Figure 1. m6A can be read by YTH-DF2, which recruits HRSP12 adapter and RNase P/MRP complex to the HRSP12-binding site and RNase P/MRP-directed cleavage site in the circRNA upstream and downstream of the YTHDF2-binding site. The circRNA is then cleaved open for rapid degradation.
Go beyond the RNA stability of m6A modification in circRNAs
m6A modified circRNAs, but not unmodified circRNAs, encode small peptides
Circular RNAs are covalently circularized RNA loops without a 5’ or 3’ end. Circular RNAs normally do not translate proteins due to the lack of a typical 5’ m7G cap structure required for protein translation initiation as in mRNAs. However, some internal m6A sites can be m6A methylated and recognized by YTHDF3 reader, which assembles protein translation initiation complex and starts protein translation (Fig.2)[13]. Hundreds of peptide products from the cap-independent translation have been confirmed by mass spectrometry.
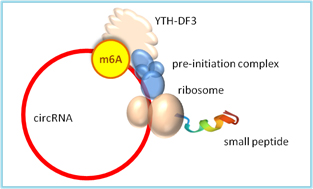
Figure 2. An m6A modification in circular RNA is sufficient for cap-independent initiation of protein translation. The m6A site is recognized by YTHDF3 reader that recruits protein translation initiation factors 4G2, 4A, 4B, and 40S small ribosomal subunit to form 43S preinitiation complex, which initiates protein translation within the RNA circle without a typical mRNA 5’-cap [13].
m6A modified circRNAs, but not unmodified circRNAs, modulate the circRNAs and RNA-binding proteins (RBP) interactions
circNSUN2 is a circular RNA frequently up-regulated in colorectal carcinoma with liver metastasis. m6A modification of circNSUN2 increases circNSUN2 export to the cytoplasm. When in the cytoplasm, the m6A modified circNSUN2 guides IGF2BP2 binding to HMGA2 mRNA and promotes the formation of circNSUN2/IGF2BP2 protein/HMGA2 mRNA ternary complex, which enhances the stability of HMGA2 mRNA and promotes HMGA2 translation(Fig.3)[14].
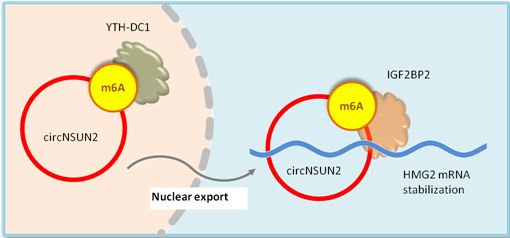
Figure 3. m6A modification of circNSUN2 increases circNSUN2 export to the cytoplasm. In the cytoplasm, m6A modified circNSUN2 promotes the circNSUN2- IGF2BP2 protein- HMGA2 mRNA ternary complex formation, which enhances HMG2 mRNA stability and translation. The increased HMG2 promotes CRC metastasis progression [14].
m6A modification of circRNAs in cancers
m6A modification of circRNAs is closely related to cancers, as circRNAs often have multiple microRNA binding sites and many microRNAs are involved in cancers as oncomiRs. As described above, m6A modified circNSUN2 ultimately leads to oncoprotein HMG2 and colorectal cancer [14].
Another example is HPV viral circular RNA circE7. m6A modification endows circE7 to translate transforming oncoprotein [15], while at the same time the m6A marks circE7 as “self” to help the virus escape from the host antiviral immune response.
m6A controls self vs foreign circRNA immunity
Foreign circRNAs (e.g. viral RNAs) can illicit antiviral gene expression (e.g. interferon) and strong immune response by the host defense. However, endogenous circRNAs, when modified by m6A and read by YTH proteins, are recognized as “self” molecules which inhibit activation of retinoic acid-inducible gene I (RIG-1) responsible for interferon production [9]. Thus m6A is served as self vs foreign circRNA marker.
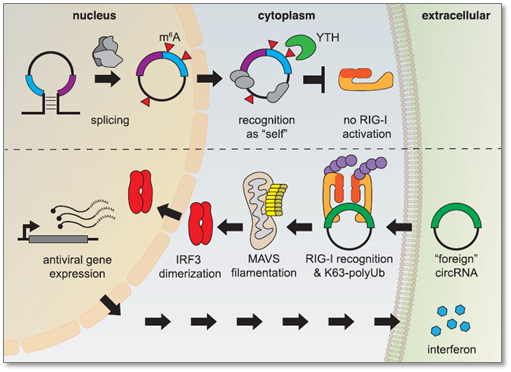
Figure 4. Endogenous circRNAs modified by m6A are recognized as “self” by YTH-DF2 reader, which block RIG-1 activation. Unmodified, foreign circRNAs activate RIG-I in the presence of K63-linked polyubiquitin, which lead to the activation of antiviral signaling pathways and interferon production [9].
With the functional roles of m6A in circRNAs not only in RNA stability, but also in small peptide translation, circRNA-miRNA sponging, circRNA-RBP binding, circRNA nuclear export, target mRNA turnover, and circular RNA immunity, profiling m6A modification of circRNAs is a new research area in the transcriptomic landscape, epitranscriptomic regulation, proteomic outcomes, and biological functions.
Related Service
CircRNA Epitranscriptomic Array Service
References
[1] Li, Z., et al. (2015) “Exon-intron circular RNAs regulate transcription in the nucleus” Nat Struct Mol Biol 22(3):256-64 [PMID: 25664725]
[2] Conn, V. M., et al. (2017) “A circRNA from SEPALLATA3 regulates splicing of its cognate mRNA through R-loop formation” Nat Plants 3:17053 [PMID: 28418376]
[3] Pamudurti, N. R., et al. (2017) “Translation of CircRNAs” Mol Cell 66(1):9-21 e7 [PMID: 28344080]
[4] Yang, Y., et al. (2018) “Novel Role of FBXW7 Circular RNA in Repressing Glioma Tumorigenesis” J Natl Cancer Inst 110(3[PMID: 28903484]
[5] Han, D., et al. (2017) “Circular RNA circMTO1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression” Hepatology 66(4):1151-1164 [PMID: 28520103]
[6] Chen, X., et al. (2018) “PRMT5 Circular RNA Promotes Metastasis of Urothelial Carcinoma of the Bladder through Sponging miR-30c to Induce Epithelial-Mesenchymal Transition” Clin Cancer Res 24(24):6319-6330 [PMID: 30305293][7] Zhou, C., et al. (2017) “Genome-Wide Maps of m6A circRNAs Identify Widespread and Cell-Type-Specific Methylation Patterns that Are Distinct from mRNAs” Cell Rep 20(9):2262-2276 [PMID: 28854373]
[8] Huang, A., et al. (2020) “Circular RNA-protein interactions: functions, mechanisms, and identification” Theranostics 10(8):3503-3517 [PMID: 32206104][9] Chen, Y. G., et al. (2019) “N6-Methyladenosine Modification Controls Circular RNA Immunity” Mol Cell [PMID: 31474572]
[10] Lewis, C. J., et al. (2017) “RNA modifications and structures cooperate to guide RNA-protein interactions” Nat Rev Mol Cell Biol 18(3):202-210 [PMID: 28144031][11] Gilbert, W. V., et al. (2016) “Messenger RNA modifications: Form, distribution, and function” Science 352(6292):1408-12 [PMID: 27313037]
[12] Park, O. H., et al. (2019) “Endoribonucleolytic Cleavage of m(6)A-Containing RNAs by RNase P/MRP Complex” Mol Cell 74(3):494-507 e8 [PMID: 30930054][13] Yang, Y., et al. (2017) “Extensive translation of circular RNAs driven by N(6)-methyladenosine” Cell Res 27(5):626-641 [PMID: 28281539]
[14] Chen, R. X., et al. (2019) “N(6)-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis” Nat Commun 10(1):4695 [PMID: 31619685]
[15] Zhao, J., et al. (2019) “Transforming activity of an oncoprotein-encoding circular RNA from human papillomavirus” Nat Commun 10(1):2300 [PMID: 31127091]
