tRNAs are thought of as abundant, ubiquitous, passive mRNA decoders and protein translators. tRNAs carrying the same amino acid are “isoacceptors”, whereas tRNAs with the same anticodon but different body sequences are “isodecoders”. Surprisingly, the latest scientific advances have found tRNA isoacceptors and isodecoders endowed with specific, non-translational, regulatory functions that can profoundly impact the biology and diseases. Additionally, tRNAs are very abundant in biofluids to be used as biomarkers. Studying these hidden functions will gain new insight into the new life of tRNAs.
| Classical textbook understanding | Latest advances |
|
|
Cell, tissue, and disease specific expression
tRNA isoacceptor and isodecoder expression profiles are tied to cell differentiation and proliferation (Fig.1A), displaying high tissue, cell type, spatial, and temporal specificity during embryonic development (Fig.1B), and disease specificity in brain, thymus, blood, spleen, liver, testis, ovary, and breast cancer (Fig.1C). The expressional specificities are associated with biological functions and pathophysiology. Also, the high specificities contribute to better tRNA biomarker performance.
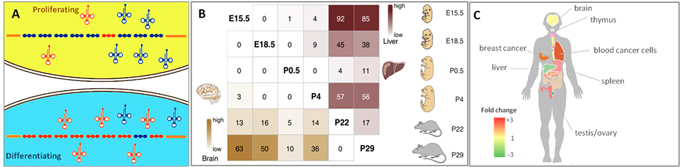
Figure 1. (A) Proliferating and differentiating cells have distinct tRNA repertoire signatures [1]. (B) tRNAs have specific expression in tissues, organs, and developmental stages [2]. (C) Differential tRNA expression in human tissues and diseases. Cancer cells generally express higher levels of nuclear encoded tRNA than every tissue examined [3].
Wobble decoding of isoacceptors affects translation
The tRNA pool, composed of tRNA isoacceptor families for each amino acid, is an important source of insider information in the study of codon usage, protein translation efficiency and accuracy, biological processes and human diseases. A codon may be decoded by a perfectly matched cognate anticodon, or, by a near-cognate tRNA with a single mismatch at the wobble position. The cognate:near-cognate tRNA ratios in the tRNA pool are dynamically regulated to affect translation efficiency, fidelity, and transcript stability (Fig. 2)[4]. Under various conditions, tRNA pools can alter in ways to maintain homeostasis or to favor a particular gene expression program [5]. The perturbed levels of isoacceptors can lead to diseases such as cancer (Fig.3A)[6] and neurodegeneration (Fig.3B)[7].
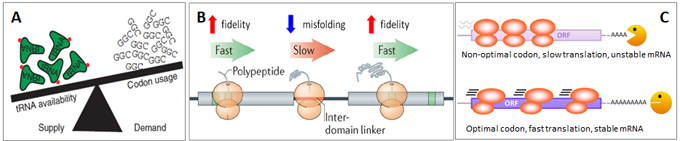
Figure 2. (A) An optimal codon is when the cognate tRNA supply is greater than the protein translation demand. A non-optimal codon is when the cognate tRNA supply is short of the demand or when near-cognate tRNAs are used. (B) Optimal codons favor faster translation speed and higher decoding accuracy. Non-optimal codons are much slower in translation but favor accurate protein folding, which tend to occur in inter-domain linker regions of the encoded proteins. (C) Optimal codons at faster translation elongation speed protect the mRNA from decay. Slow non-optimal codons expose the mRNA for degradation and shorten the mRNA half-lives.
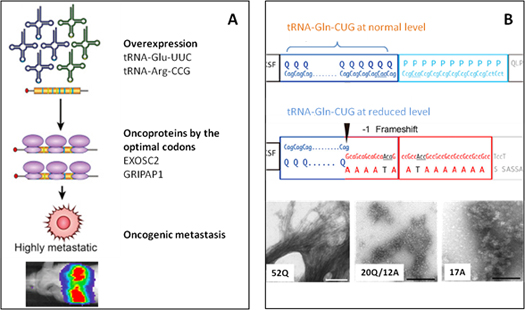
Figure 3. (A) Overexpression of tRNA-Glu-UUC and tRNA-Arg-CCG is sufficient to drive oncogenic metastasis with their optimal codons up-regulating oncoprotein EXOSC2 and GRIPAP1 translation. [6]. (B) tRNA-Gln-CUG at normal supply level correctly translates poly-Q repeat region in Huntintin. Reduced tRNA-Gln-CUG level causes -1 frame shift translation to make erroneous poly-Alanines. The extent of Q/A ratio in Huntingtin determines the aggregation state (from 52Q normal, 20Q/12A intermediate, to 17A pathogenic) and thus the severity of Huntington’s disease [7].
Isodecoders impact on biological processes and diseases
The highly conserved tRNA isodecoder sequences can exhibit different tissue-specific expression and perform significantly different functions. For example, one of five mouse tRNA-Arg-UCU isodecoders is a central nervous system (CNS) specific isodecoder (Fig.4A). Its deficiency can cause ribosome stalling on mRNAs, ATF4-driven stress response, and wide spread neurodegeneration (Fig.4B, C, D). Thus, the CNS-specific isodecoder has a new function of maintaining neuronal cell homeostasis and preventing neurodegeneration [8].
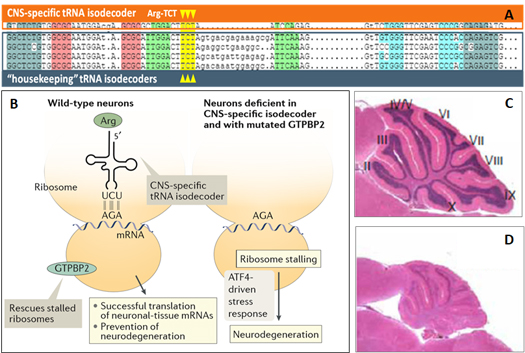
Figure 4. (A) CNS-specific and “housekeeping” isodecoder genes of tRNA-Arg-UCU. (B) CNS-specific isodecoder is vital for successful translation in wild type neurons, without which the cells undergo neurodegeneration. (C) Normal wild type brain. (D) Neurodegenerative brain in the absence of CNS-specific tRNA-Arg-UCU isodecoder [8].
Non-canonical, non-translational, regulatory tRNA functions
tRNAs can regulate cellular functions not by direct protein translation. For example, tRNAs can bind and inhibit cytochrome C from forming apoptosome in apoptosis [9,10]. Thus, tRNAs are an anti-apoptosis factor.
tRNAs are also the first responder to cellular stress such as heat shock, hypoxic shock or nutrient deprivation by multiple regulatory mechanisms to shut down global protein translation (Fig. 5)[11].
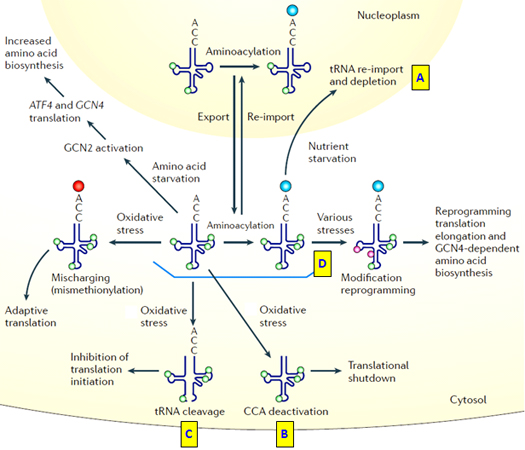
Figure 5. tRNAs in response to cellular stresses by (A) Re-importation of tRNAs from cytoplasm to the nucleus, depleting tRNA pool in cytosol for protein translation; (B) Deactivation of tRNA 3’CCA arm; (C) Hypoxia induced tRNA cleavage to form tRF and tiRNA fragments that inhibit protein translation initiation. (D) During nutrient starvation, re-programming tRNA modifications to favor translation of proteins/enzymes in amino acid biosynthesis pathway [11].
tRNA biomarkers in biofluids
Small RNAs such as microRNAs have been popularly explored as biomarkers. tRNAs are now emerging as a new class of biomarkers, owing to their many desired characteristics. They are abundant small RNAs and particularly abundant in biofluids, even more so than miRNAs (Fig. 6A)[12, 13]. tRNAs are dramatically enriched in exosomes to become the most abundant RNA class second only to rRNAs, far more than miRNAs (Fig. 6B)[14].
The high abundance, stability protected by heavy modifications, and enrichment in biofluids mean better biomarker sensitivity. Together with the tight association of perturbed tRNA repertoires with diseases (Fig. 1), tRNAs can be an class of biomarkers with greater potentials.
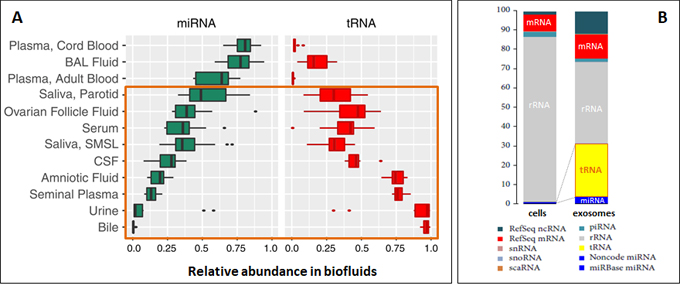
Figure 6. (A) Relative proportions of miRNA vs tRNA in biofluids, where many biofluids have tRNA contents much higher than miRNA [12, 13]. (B) tRNAs are highly enriched in exosomal RNA compared with intracellular RNA, much more so than miRNAs [14].
Related Product
References
[1] Gingold H. et al. (2014) Cell [PMID: 25215487]
[2] Schmitt B.M. et al. (2014) Genome Res. [PMID: 25122613]
[3] Geslain R. and G. Eriani (2014) Translation [PMID: 26779404]
[4] Hanson G. and J. Coller (2018) Nat. Rev. Mol. Cell Biol. [PMID: 29018283]
[5] Gavin H. et al. (2014) Molecular Cell Biology [PMID: 29018283]
[6] Goodarzi H. et al. (2016) Cell [PMID: 27259150]
[7] Girstmair H. et al. (2013) Cell Rep [PMID: 23352662]
[8] Ishimura R. et al. (2014) Science [PMID: 25061210]
[9] Mei Y. et al. (2010) Mol. Cell [PMID: 20227371]
[10] Mei Y. et al. (2010) Protein Cell [PMID: 21113408]
[11] Kirchner S. and Z. Ignatova (2015) Nat. Rev. Genet. [PMID: 25534324]
[12] Godoy P.M. et al. (2018) Cell Rep [PMID: 30380423]
[13] Schageman J. et al. (2013) Biomed Res Int [PMID: 24205503]
[14] Quek C. et al. (2017) RNA Biol [PMID: 28005467]
