Super-enhancer lncRNAs have been demonstrated master regulatory functions in biology and diseases, such as myogenesis, cardiomyocyte differentiation, erythropoiesis, fibrosis in post-myocardial infarction, and cancers. The innovative Arraystar Super-enhancer LncRNA Arrays simultaneously profile the super-enhancer lncRNAs and their target genes. The detailed annotation, analysis and cross-references are provided to facilitate the in-depth study on SE-lncRNAs and understanding the biological functions.
Nearby target gene annotation
The most common mechanism of SE-lncRNA regulation is by cis-activation, as in the examples shown in Fig. 1. Hand2, a key transcription factor controlling cell differentiation from fibroblasts to cardiomyocytes, is regulated by the nearby SE-lncRNA upperhand (Uph) [1] (Fig.1A); SE-lncRNA CCAT1-L produced by the super-enhancer upstream of MYC oncogene forms the enhancer:promoter chromatin loops to stimulate MYC transcription and colorectal cancer [2](Fig.1B); The SE-lncRNA CERNA of the MyoD super-enhancer can recruit activating chromatin modifier and load RNA polymerase II to the MyoD promoter [3] (Fig.1C, Fig.2). Arraystar furnishes the information for the nearby genes overlapping with or within 50-kb distance of the SE-lncRNAs, which helps to quickly locate the potential target genes under the SE-lncRNA regulation.
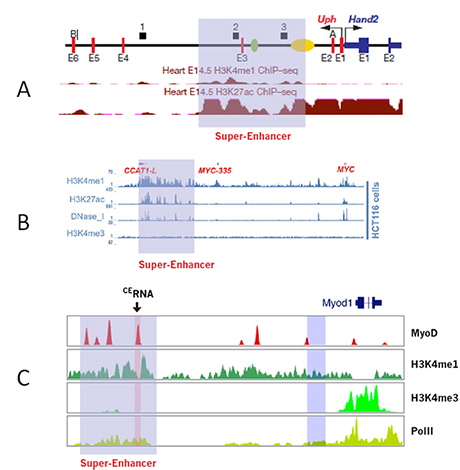
Figure 1. Examples of SE-lncRNAs and their nearby genes. (A) SE-lncRNA upperhand (Uph) and its target gene Hand2, a key transcription factor in cardiomyocyte differentiation [1]. (B) SE-lncRNA CCAT1-L and its target gene MYC, an oncogene in colorectal cancer [2]. (C) SE-lncRNA Core Enhancer RNA (CERNA) and its target gene MyoD, a master transcription factor in myogenesis [3].
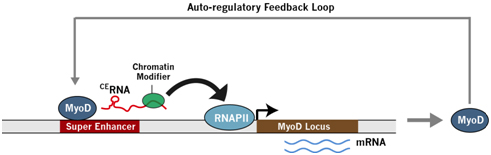
Figure 2. SE-lncRNA regulating the nearby gene expression. MyoD is the master transcription factor in myogenesis, controlling other key transcription factors during the myogenic gene expression program. Upon myogenic stimulation, MyoD,the core enhancer (CE) in the super-enhancer upstream of the MyoD gene transcribes a SE-lncRNA CERNA. CERNA recruits activating chromatin modifier to open up the chromatin and also increases the RNA polymerase II loading to the MyoD promoter, leading to MyoD upregulation. MyoD further activates its own CERNA, forming a positive auto-regulatory feedback loop [3, 4].
Super-lncRNA
“Super-lncRNAs”, a special class of super-enhancer lncRNA, form RNA:DNA:DNA triplex structure with the host super-enhancers by Hoogsteen and reverse Hoogsteen base pairing. Super lncRNAs can recruit transcription factors to the super-enhancers, alter the chromatin structure, act as spatial amplifiers, and increase the tissue specificity of the gene expression associated with the super-enhancers. As the majority of the super lncRNAs are transcribed in the super-enhancer regions, the annotation of super lncRNAs lends the research rationales toward this class of SE-lncRNAs.
Subcellular localization
The functional mechanisms long non-coding RNAs (lncRNAs) are tightly coupled with their subcellular localization. For example, lncRNAs localized in the nucleus or chromatin often regulate the gene expression by epigenetic modification and transcription. LncRNAs in the cytoplasm are more likely involved in translation regulation or miRNA sponging such as competing endogenous RNAs (ceRNA). SE-lncRNAs have increasingly been found to act in trans. Therefore, the subcellular localization information provides an important clue in knowing SE-lncRNA functional mechanism.
Tissue or cell type specificity
Super-enhancer activities exhibit high degrees of tissue or temporal specificity, which are tightly associated with the very genes that control the tissue specificity. The transcription of SE-lncRNAs is the hallmark of the super-enhancers in an activated status. Furthermore, SE-lncRNAs cooperate with the host super-enhancers to accomplish gene regulation by multiple mechanisms. The cell lineage specificities of the SE-lncRNAs are very informative to gain insight into the functional mechanisms of the specificity.
Cancer and disease SE-lncRNAs
SE-lncRNAs are associated with human diseases and particularly with cancers. The cancer SE-lncRNAs and diseases SE-lncRNAs are annotated with databases such as Lnc2Cancer database and lncRNADisease.
Related Services
References
1. Anderson KM et al: Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development. Nature 2016, 539(7629):433-436.[PMID: 27783597]
2. Xiang JF et al: Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res 2014, 24(5):513-531.[PMID: 24662484]
3. Mousavi K et al: eRNAs promote transcription by establishing chromatin accessibility at defined genomic loci. Mol Cell 2013, 51(5):606-617.[PMID: 23993744]
4. Shiekhattar R: Opening the Chromatin by eRNAs. Mol Cell 2013, 51(5):557-558.[PMID: 24034693]
