Related Products
Related Services
Related Reviews
NuRNA™ tRNA Modification Enzymes PCR Array (H)
Benefits
• Best content –covers all the tRNA modification enzymes in UniProt and Modomics
• Rigorous – all primer pairs and assays are elaborately optimized and rigorously validated
• Convenient – easy-to-use plate format for direct sample application for full profiling. No sample pre-amplification is needed.
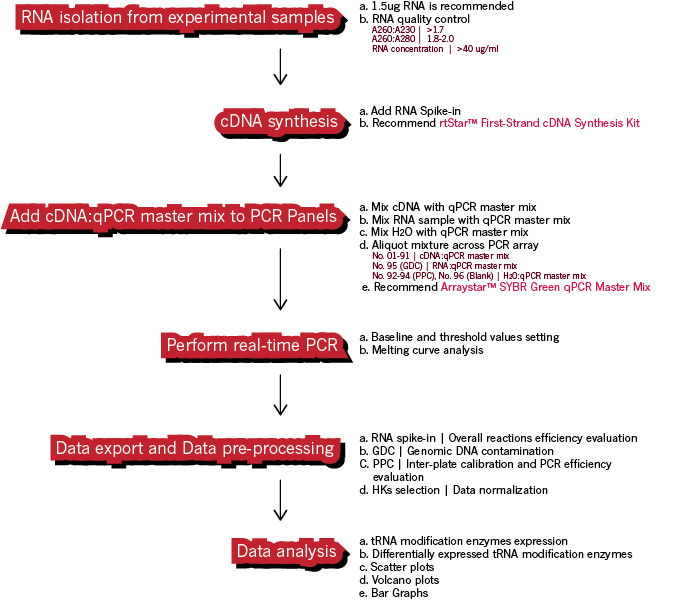
We recommend using our rtStar™ First-Strand cDNA Synthesis Kit (AS-FS-001) along with Arraystar SYBR® Green Real-time qPCR Master Mix when running your NuRNA™ tRNA Modification Enzymes PCR Array for optimal performance.
| Product Name | Catalog No. | Description | Size | Price |
|---|---|---|---|---|
| NuRNA™ Human tRNA Modification Enzymes PCR Array | AS-NM-001-1 | 384 (4x96)-well plate | $567.00 | |
| NuRNA™ Human tRNA Modification Enzymes PCR Array (Roche Light Cycler 480) | AS-NM-001-1-R | 384 (4x96)-well plate | $567.00 |
tRNAs are the key component in protein translation to decode genetic codes with the charged amino acids. Recent studies have revealed tRNAs, as well as their derived small noncoding RNA fragments, are dynamically regulated and have non-canonical functions such as adaptive translation, cell proliferation/differentiation, stress response, metabolism, and diseases[1].
tRNAs are heavily decorated posttranscriptionally with numerous chemical modifications. The modifications are essential for shaping up, fine tuning, and regulating all aspects of tRNA functioning, such as folding, stability and decoding. For example, modifications in the stem-loops are crucial for tRNA structure and stability, in anticodon loop for translation accuracy, at position 34 for codon-anticodon wobbling, and adjacent to anticodon loop for fine tuning the codon-anticodon interaction (Fig.1).
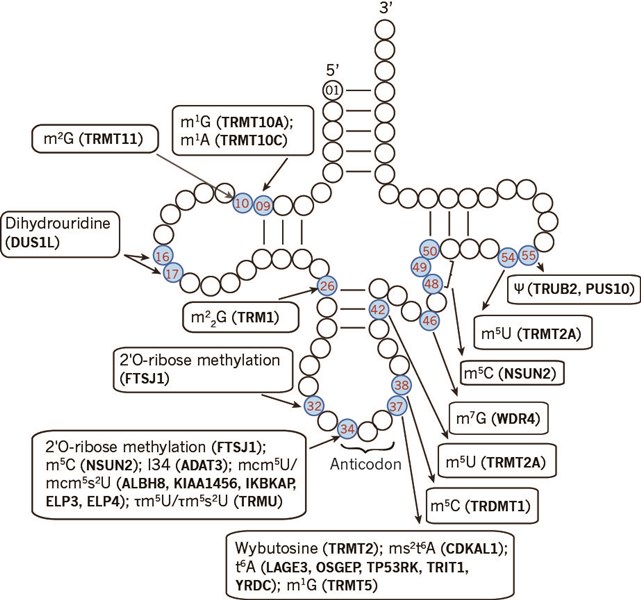
Figure 1. Human tRNA modifications and tRNA modification enzymes (in parentheses).
These chemical modifications are dynamically regulated and catalyzed by tRNA modification enzymes[2]. Mutations or dysregulation of tRNA modifiers have been associated with diseases, for example, mutations in cytosine-5 RNA methyltransferase (NSUN2) with microcephaly[3] and t6A methylthiolation enzymes with type 2 diabetes[4]. The emerging importance of tRNA modifications in diseases calls for additional work[5, 6]. How tRNA modifications are regulated by tRNA modification enzymes is in need to be studied.
To help researchers quickly and conveniently profile enzymes involved in tRNA modifications, Arraystar has produced the first commercial PCR panel specially designed for profiling the tRNA modification enzymes. The panel contains 85 validated or predicted tRNA modification enzymes or protein factors compiled from research publications and authoritative databases including UniProt and Modomics. Each assay on the panel is rigorously validated across many tissues and cell lines. Spike-in control, Positive PCR Control, Genomic DNA Control and normalization references are included to ensure the upmost data quality.
References
[1] Kirchner S, Ignatova Z. Emerging roles of tRNA in adaptive translation, signalling dynamics and disease. Nature reviews Genetics 2015;16:98-112.
[2] El Yacoubi B, Bailly M, de Crecy-Lagard V. Biosynthesis and function of posttranscriptional modifications of transfer RNAs. Annual review of genetics 2012;46:69-95.
[3] Blanco S, Dietmann S, Flores JV, Hussain S, Kutter C, Humphreys P, et al. Aberrant methylation of tRNAs links cellular stress to neuro-developmental disorders. The EMBO journal 2014;33:2020-39.
[4] Zhou B, Wei FY, Kanai N, Fujimura A, Kaitsuka T, Tomizawa K. Identification of a splicing variant that regulates type 2 diabetes risk factor CDKAL1 level by a coding-independent mechanism in human. Human molecular genetics 2014;23:4639-50.
[5] Zhang X, Cozen AE, Liu Y, Chen Q, Lowe TM. Small RNA Modifications: Integral to Function and Disease. Trends in molecular medicine 2016.
[6] Suzuki T, Nagao A, Suzuki T. Human mitochondrial tRNAs: biogenesis, function, structural aspects, and diseases. Annual review of genetics 2011;45:299-329.
| 85 Human tRNA Modification Enzymes/Proteins and the Modifications (in parenthesis) |
| ADAT1(m1I37), ADAT2(m1I34), ADAT3(-), ALKBH1(mcm5U), ALKBH8(mchm5U34/mcm5s2U34/mcm5U34/mcm5Um34), C9orf64(-), CDK5RAP1(ms2A37/ms2 i6A37), CDKAL1(ms2A37/ms2 i6A37), CDKL1(ms2t6A), CTU1(mcm5S2U ), CTU2(mcm5S2U), DUS1L(D16/17), DUS2(D20), DUS3L(D47), DUS4L(D), ELP3(-), ELP4(mcm5s2U), FBLL1(-), FTSJ1(Cm32/Gm34), GTPBP3(tm5U34), HSD17B10(m1A9/m1G9), IKBKAP(-), KIAA0391(m1A9/m1G9), KIAA1456(Um), LAGE3(t6A37), LCMT2(o2yW), METTL1(m7G46), METTL2A(Cm), METTL2B(3mC), MOCS3(mcm5S2U ), MTO1(tm5U34), NAT10(Ac4C), NFS1(s2U34/tm5s2U34), NSUN2(m5C49), NSUN6(m), OSGEP(t6A37), OSGEPL1(t6A37), PUS1(pseudouridine27/pseudouridine28), PUS10(pseudouridine 55), PUS3(pseudouridine39), PUS7L(pseudouridine), PUSL1(pseudouridine), QTRT1 (Q34), QTRT2 (Q34), RPUSD1(pseudouridine), RPUSD2(pseudouridine31/pseudouridine32), RPUSD3(pseudouridine), RPUSD4(pseudouridine31/pseudouridine32), SSB(-), TARBP1(-), THUMPD1(Acetylcytidine), THUMPD2(Gm), THUMPD3(Gm), TP53RK(t6A37), TPRKB(t6A37), TRDMT1(m5C38), TRIT1(i6A37), TRMO(m6t6A), TRMT1(m2G26/m22G26), TRMT10A(m1G9), TRMT10B(m1G9), TRMT10C(m1A9/m1G9), TRMT11(m2G10), TRMT112(mchm5U34/mcm5s2U34/mcm5U34/mcm5Um34), TRMT12(o2yW), TRMT13(m4C/m4A ), TRMT1L(m2G26/m22G26), TRMT2A(Um), TRMT2B(m5U54), TRMT44(Um44), TRMT5(m1G37/o2yW), TRMT6(m1A58), TRMT61A(m1A58), TRMT61B(m1A58), TRMU(s2U34/tm5s2U34), TRUB1(pseudouridine), TRUB2(pseudouridine55), TYW1(o2yW), TYW1B(o2yW), TYW3(o2yW), TYW5 (o2yW), UBA5(cyclic t6A), URM1(mcm5S2U), WDR4(m7G46), YRDC(t6A37) |
Compatible qPCR Instruments Equipped with a 384-well Format Block:
ABI ViiA™ 7
ABI 7500 & ABI 7500 FAST
ABI 7900HT
ABI QuantStudio™ 5 Real-Time PCR system
ABI QuantStudio™ 6 Flex Real-Time PCR system
ABI QuantStudio™ 7 Flex Real-Time PCR system
ABI QuantStudio™ 12K Flex Real-Time PCR System
Bio-Rad CFX384
Bio-Rad iCycler & iQ Real-Time PCR Systems
Eppendorf Realplex
Stratagene Mx3000
Roche Light Cycler 480

